种质资源是农业生产的物质基础,是科技创新的源头。中国是种质资源大国,但相关的发掘与利用工作却存在严重不足,这与基因型鉴定不系统、不全面有一定关系。基因型是指某一生物个体全部基因组合的总称,而遗传学中往往特指某一性状的基因型。随着高通量测序技术的出现,一大批关键基因得到发掘,基因型鉴定工作也随之取得一定进展。曹云[1]选择了来自全国不同生态区的10个小麦代表品种,采用分子标记技术确定了各品种的春化与光周期发育基因型,强春性春播春小麦为Vrn1三个位点全显性或Vrn-A1显性,春性或弱春性秋播冬小麦为Vrn-B1显性或Vrn-D1显性,冬性半冬性冬小麦为Vrn1三个位点全隐性。在栽培花生上,油酸含量由2个同源非等位基因AhFAD2A和AhFAD2B共同调控。研究者对16个花生品种进行鉴定,发现低油酸品种(系)的基因型均为AABB,中油酸品种(系)的基因型为aaBB,高油酸品种(系)的基因型为aabb[2]。在果树上,赵雅楠等[3]、刘春生等[4]和张号楠等[5]利用蔷薇科S-RNɑse通用引物对20份中国李、60份苹果和39份甜樱桃材料进行了S基因型鉴定,为这些树种的授粉树配置提供了理论支撑。王朝晖等[6]鉴定189份桃种质与肉质形成相关的F-M基因座、YUC11启动子区转座子插入、NAC72 编码区9 bp插入的基因型,明确了不同种质在3个位点的基因型组合形式。吉爽秋等[7]利用关联分析,发现在桃第8染色体的14 484 624 bp处存在1个与花型关联的单碱基核苷酸变异(SNP),当其基因型为CC、AC时,其表型为铃形;基因型为AA时,表型为蔷薇形,在145份桃自然群体中鉴定准确率为98.62%。
在上述研究中,基因型鉴定结果通常针对质量性状,能够反映出表型与各单倍型之间的关系,却较少考虑同一性状受到不同调控基因的影响。肖鑫等[8]利用特异性引物通过PCR技术对72个酿酒葡萄品种的MYBA1和MYBA2基因位点的基因型和单倍型进行鉴定,发现酿酒葡萄2个MYB基因位点共有10个等位基因,其组成的不同单倍型参与调控果实着色,虽然单倍型数量与果皮颜色深浅没有直接的关系,但总体上呈现出着色功能基因越多则果皮颜色越深的规律。
桃是中国重要的果树种类,栽培面积居落叶果树第3位,在促进农民增收和乡村振兴方面发挥着重要作用。在桃上,目前已经发掘出的关键基因主要包括果实肉色白/黄性状基因PpcCCD4[9]、柱形性状基因PpeTAC1[10]、果皮茸毛性状基因Ppe-MYB25[11]、果实硬质性状基因PpYUC11[12]、果实溶质性状基因PpendoPGM[13]、矮化性状基因Ppe-GID1c[14]、垂枝性状基因ppɑ013325m[15]、蟠桃果形性状基因PpOFP1[16]、花色非白/白性状基因PpGST[17]等以及果实糖含量[18-20]、酸含量[21]、果肉花色素苷含量[22]和其他一些抗性性状如抗蚜[23]和抗根结线虫[24]等关键基因。单果质量(旧称单果重)是果树作物长期以来重要的育种目标之一,因此,不少研究针对桃单果质量性状进行了连锁分析工作,在桃的8个连锁群(染色体)上均有单果质量QTLs定位[25-31]。此外,利用全基因组关联分析(GWAS),鉴定到一些单果质量关联位点。Cao等[32]利用129份桃种质重测序鉴定出的4 063 377个SNPs,对2007年和2010年的单果质量表型进行了GWAS分析,在桃的8条染色体上均鉴定出关联位点。随后,Cao等[33]利用更大的群体(313份桃种质)进行重测序,对该自然群体连续3年的单果质量性状进行了GWAS分析,在第2和第6染色体上共鉴定出5个关联位点。Ksouri等[34]利用90份桃种质鉴定出的13 045个高质量SNPs对单果质量进行GWAS分析,发现显著的关联位点在第3、6、8染色体上,表型变异解释率介于17%~22%之间。Wang等[35]提出Fast3VmrMLM算法,通过整合全基因组扫描与机器学习,可高效鉴定SNP标记、基因单倍型等与数量性状的关联,在水稻、玉米中成功挖掘出大量相关基因。Zhang等[36]开发葡萄200K Axiom SNP芯片(标记密度每2.8 kb一个,分辨率提升10倍),结合高通量表型平台(每秒获取单果124个性状)及复合性状降维算法,定位到调控果形的bHLH017基因和抗寒关键基因NAC08。Wang等[37]利用稻瘟病抗性基因Pi-tɑ和Pi-b的分子标记,对育种群体进行检测并聚合抗病基因,结合快速加代与抗性选择,选育出高产多抗的节水抗旱稻新品种沪旱106,为数量性状定向改良提供了参考。在桃上关于分子育种的策略研究报道较少。前人对桃果实单果质量性状进行了不少的连锁和关联分析,鉴定出的分子标记对理解该性状形成的遗传机制以及分子标记辅助育种具有重要意义。然而,目前的研究多倾向于发掘优异基因及连锁/关联标记,对不同种质的单个基因或标记的基因型鉴定较少,对不同标记对单果质量表型的贡献分析也较少。本研究在总结前人鉴定的单果质量分子标记基础上,结合574份种质基因组重测序数据,在只考虑位点间加性作用的情况下,分析中国育成品种的单果质量育种值,筛选推荐可用于改良现有品种的亲本材料,为中国桃产业健康发展提供重要的理论和技术支撑。
1 材料和方法
1.1 材料
供试材料为574份桃种质资源,及中油蟠7号、中油蟠15号和华光3个育种品种,均种植于中国农业科学院郑州果树研究所国家果树种质郑州桃圃和中国农业科学院郑州果树研究所桃种质资源团队试验基地(新乡),树势生长良好,无病虫害危害,处于稳定结果期,田间管理措施一致。
1.2 单果质量评价
574份种质的单果质量表型来自《中国桃遗传资源》[38]。另外3份系谱关系清晰的育成品种的单果质量表型主要来自品种审定时的平均单果质量。
1.3 基因型鉴定
574份种质的基因型数据来自实验室前期全基因组重测序,已提交至基因组序列数据库NCBI,登录号为PRJCA025647。利用Trimmomatic软件[39]对原始测序数据进行质控,获得过滤后的序列。随后使用BWA软件[40]进行比对,进一步使用GATK软件[41]进行变异检测,最后获得每个品种的基因型。
1.4 基因型对单果质量性状的效应值分析
参考Ksouri等[34]的方法计算基因型的效应值,通过成对比较不同基因型(纯合参考0/0、杂合0/1和纯合突变位点1/1)的表型值,估计SNP位点对性状的等位基因效应。当观察到纯合参考和其余基因型之间表型值相比为显著增加时,等位基因被定义为正向;否则定义为反向。
2 结果与分析
2.1 单果质量性状连锁/关联标记筛选
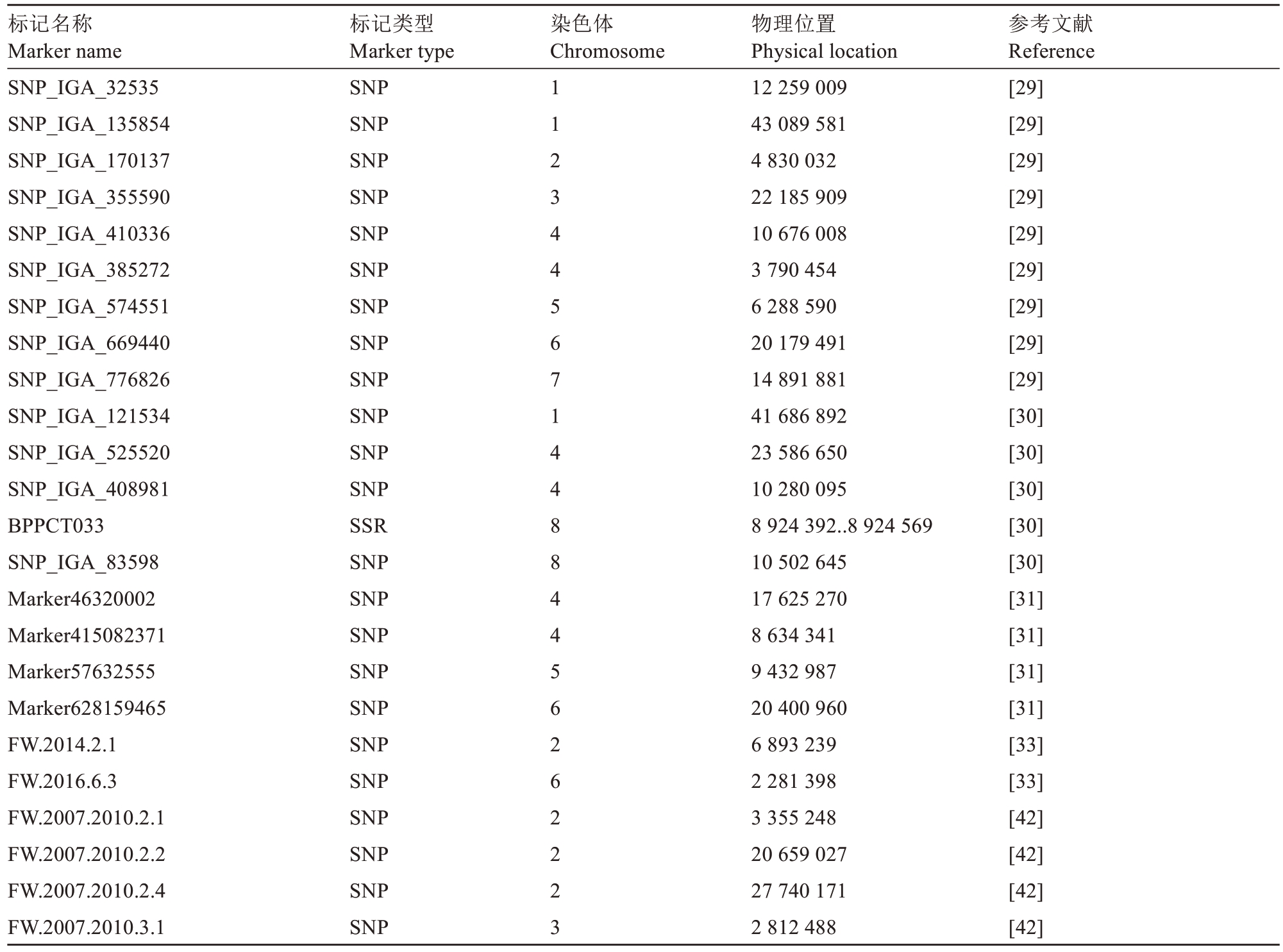
参考国内外研究进展,也为便于从基因组重测序数据中获得不同种质的基因型信息,本研究选择了SNP、Indel等有特定染色体位置的46个标记,这些标记来自8个研究团队。第1到第8染色体均有标记分布,数目分别为5、6、4、9、3、12、2和5个,即定位在第4和6染色体的位点较多。在第4染色体上的9个标记位点中,有3个都定位在10.2~10.7 Mb之间;在第6染色体上的12个标记位点中,有4个都定位在1.8~2.3 Mb之间。上述两个区段在不同研究者间具有较高的重复性。具体的单果质量连锁/关联标记信息如表1所示。
表1 单果质量连锁/关联标记信息
Table 1 The linkage/association marker information of fruit weight
标记名称Marker name标记类型Marker type染色体Chromosome物理位置Physical location参考文献Reference SNP_IGA_32535 SNP_IGA_135854 SNP_IGA_170137 SNP_IGA_355590 SNP_IGA_410336 SNP_IGA_385272 SNP_IGA_574551 SNP_IGA_669440 SNP_IGA_776826 SNP_IGA_121534 SNP_IGA_525520 SNP_IGA_408981 BPPCT033 SNP_IGA_83598 Marker46320002 Marker415082371 Marker57632555 Marker628159465 FW.2014.2.1 FW.2016.6.3 FW.2007.2010.2.1 FW.2007.2010.2.2 FW.2007.2010.2.4 FW.2007.2010.3.1 SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SSR SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP 1 1 2 3 4 4 5 6 7 1 4 4 8 8 4 4 5 6 2 6 2 2 2 3 12 259 009 43 089 581 4 830 032 22 185 909 10 676 008 3 790 454 6 288 590 20 179 491 14 891 881 41 686 892 23 586 650 10 280 095 8 924 392..8 924 569 10 502 645 17 625 270 8 634 341 9 432 987 20 400 960 6 893 239 2 281 398 3 355 248 20 659 027 27 740 171 2 812 488[29] [29] [29] [29] [29] [29] [29] [29] [29] [30] [30] [30] [30] [30] [31] [31] [31] [31] [33] [33] [42] [42] [42] [42]
表1(续) Table 1 (Continued)
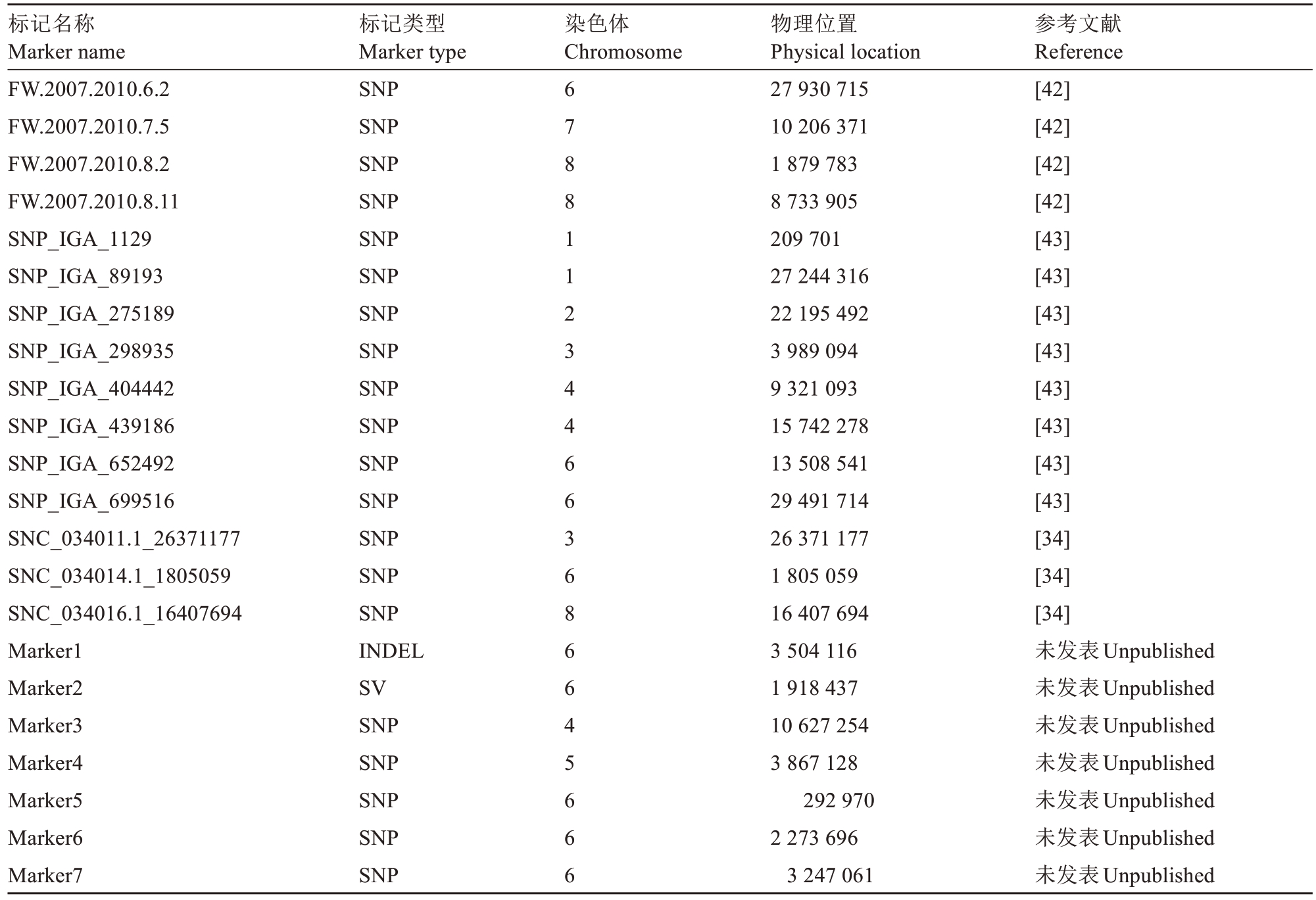
注:物理位置指的是该位点在基因组V2.0上的位置。“未发表”指的是本团队正在进行关联分析或连锁分析得到的桃单果质量定位信息。
Note:The physical location indicates where this locus is situated on genome version 2.0. ‘Unpublished’ refers to the positioning data of peach fruit weight acquired by our team viɑ correlation or linkage analysis.
标记名称Marker name标记类型Marker type染色体Chromosome物理位置Physical location参考文献Reference FW.2007.2010.6.2 FW.2007.2010.7.5 FW.2007.2010.8.2 FW.2007.2010.8.11 SNP_IGA_1129 SNP_IGA_89193 SNP_IGA_275189 SNP_IGA_298935 SNP_IGA_404442 SNP_IGA_439186 SNP_IGA_652492 SNP_IGA_699516 SNC_034011.1_26371177 SNC_034014.1_1805059 SNC_034016.1_16407694 Marker1 Marker2 Marker3 Marker4 Marker5 Marker6 Marker7 SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP SNP INDEL SV SNP SNP SNP SNP SNP 6 7 8 8 1 1 2 3 4 4 6 6 3 6 8 6 6 4 5 6 6 6 27 930 715 10 206 371 1 879 783 8 733 905 209 701 27 244 316 22 195 492 3 989 094 9 321 093 15 742 278 13 508 541 29 491 714 26 371 177 1 805 059 16 407 694 3 504 116 1 918 437 10 627 254 3 867 128 292 970 2 273 696 3 247 061[42] [42] [42] [42] [43] [43] [43] [43] [43] [43] [43] [43] [34] [34] [34] 未发表Unpublished未发表Unpublished未发表Unpublished未发表Unpublished未发表Unpublished未发表Unpublished未发表Unpublished
2.2 单果质量性状连锁/关联标记过滤及对单果质量性状的效应值分析
为进一步合并表1中的位点,利用实验室前期的574份桃栽培品种重测序的遗传变异信息,计算了种质连锁不平衡(linkage disequilibrium,LD),用LD的强度(r2)衰减至一半时的物理距离(LD decay)作为位点合并的阈值。分析表明,桃LD decay值为10.52 kb(图1),即当不同研究间的位点距离小于10.52 kb时,仅考虑一个位点(优先考虑具有功能验证证据的位点),如表1中的FW.2016.6.3和Marker6。经过合并,位点缩减至25个;进一步分析发现,有6个在574份种质中严重偏分离,因此最终确定19个位点可以用来计算其在自然群体中对单果质量性状形成所贡献的效应值。
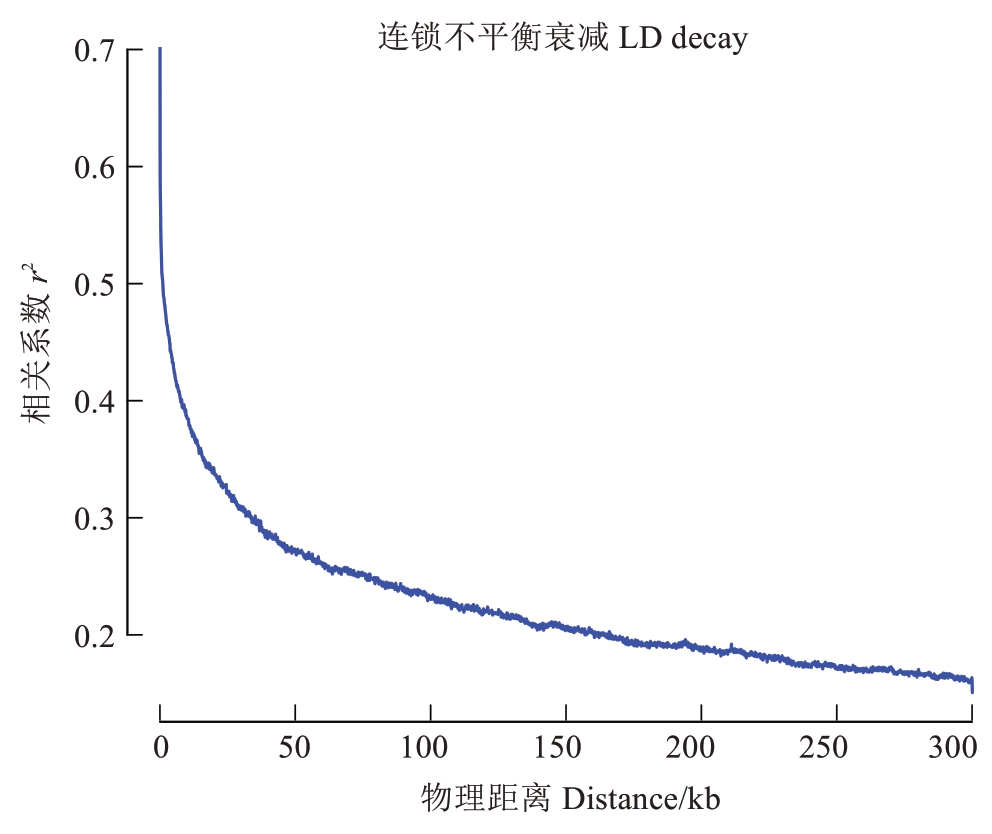
图1 全基因组连锁不平衡衰减距离的计算
Fig. 1 Calculation of physical distance for genome-wide linkage disequilibrium
分析结果显示(表2),19个位点的效应值呈现显著异质性(范围:-20.06~10.86)。其中3个位点(N3、N10和N19)表现出强的正向效应,位于第1染色体末端的N3效应值最高,达到10.86;而5个位点(N11、N13、N15、N16和N17)具有强的负向效应,位于第6染色体顶端的N15效应值最低,达到-20.06;中性效应位点(-1.72~1.04)占比36.84%,表明该性状受多个微效基因控制,且负向效应位点对表型变异的贡献更为突出。
表2 19个位点对单果质量性状形成贡献的效应值
Table 2 Effect values of 19 loci on the formation of fruit weight traits
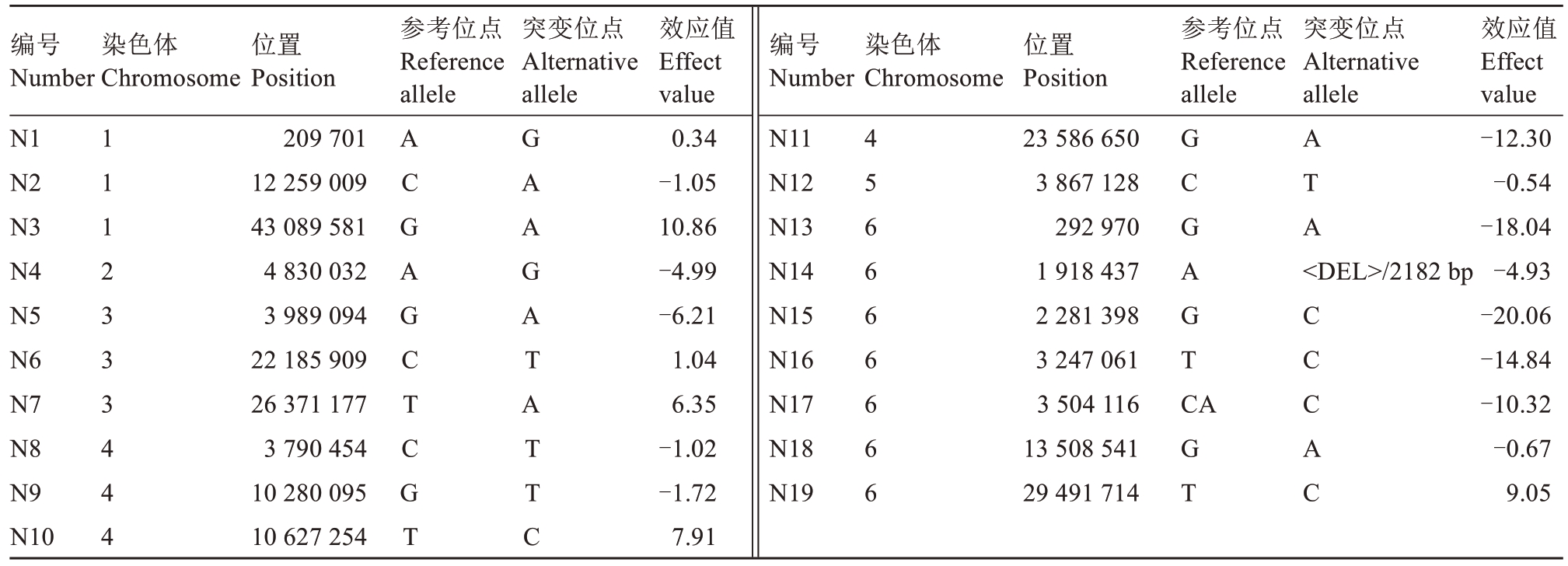
注:正负符号反映该等位基因对单果质量性状的影响方向,正值表示该等位基因具有增加单果质量性状的效应,负值表示该等位基因具有降低单果质量性状的效应。
Note:The positive and negative signs indicates the direction of the influence of the allele on the fruit weight trait. Positive values indicate that the allele has the effect of increasing the fruit weight,while negative values indicate that the allele has the effect of decreasing the fruit weight.
编号Number N1 N2 N3 N4 N5 N6 N7 N8 N9 N10染色体Chromosome参考位点Reference allele突变位点Alternative allele染色体Chromosome参考位点Reference allele突变位点Alternative allele 1 1 1 2 3 3 3 4 4 4位置Position 209 701 12 259 009 43 089 581 4 830 032 3 989 094 22 185 909 26 371 177 3 790 454 10 280 095 10 627 254 A C G A G C T C G T G A A G A T A T T C效应值Effect value 0.34-1.05 10.86-4.99-6.21 1.04 6.35-1.02-1.72 7.91编号Number N11 N12 N13 N14 N15 N16 N17 N18 N19 4 5 6 6 6 6 6 6 6位置Position 23 586 650 3 867 128 292 970 1 918 437 2 281 398 3 247 061 3 504 116 13 508 541 29 491 714 G C G A G T C A T A/2182 bp A G T C C C A C效应值Effect value-12.30-0.54-18.04-4.93-20.06-14.84-10.32-0.67 9.05
2.3 3个优良品种育种过程中亲本材料的基因型效应解析
为验证不同位点效应值的可靠性,本研究基于筛选获得的19个单果质量相关分子标记和重测序数据,对中油蟠7号(300 g)(图2)、中油蟠15号(200 g)(图3)及华光(78 g)(图4)的育种进程进行系统追踪,分析其亲本材料的单果质量育种值的动态变化。分析发现,中油蟠7号和中油蟠15号在选育过程中,亲本材料的育种值略有起伏,但整体呈现持续增大趋势;华光在育成过程中,单果质量的育种值呈现直线下降趋势,与其选育过程中实际单果质量的下降趋势一致。
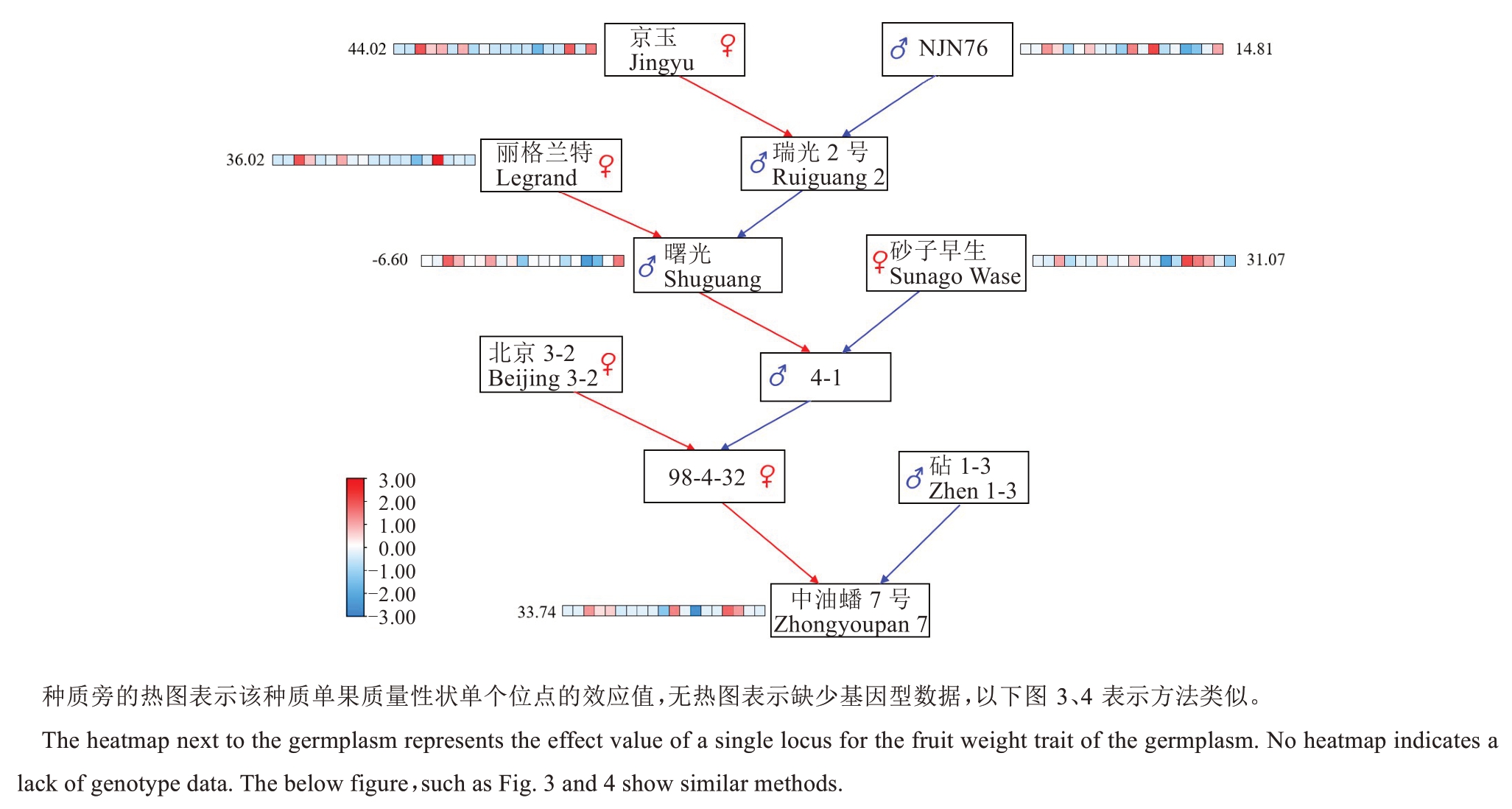
图2 中油蟠7号选育过程中亲本材料育种值的变化Fig. 2 Changes in breeding values of parental materials during the breeding process of Zhongyoupan 7 peach
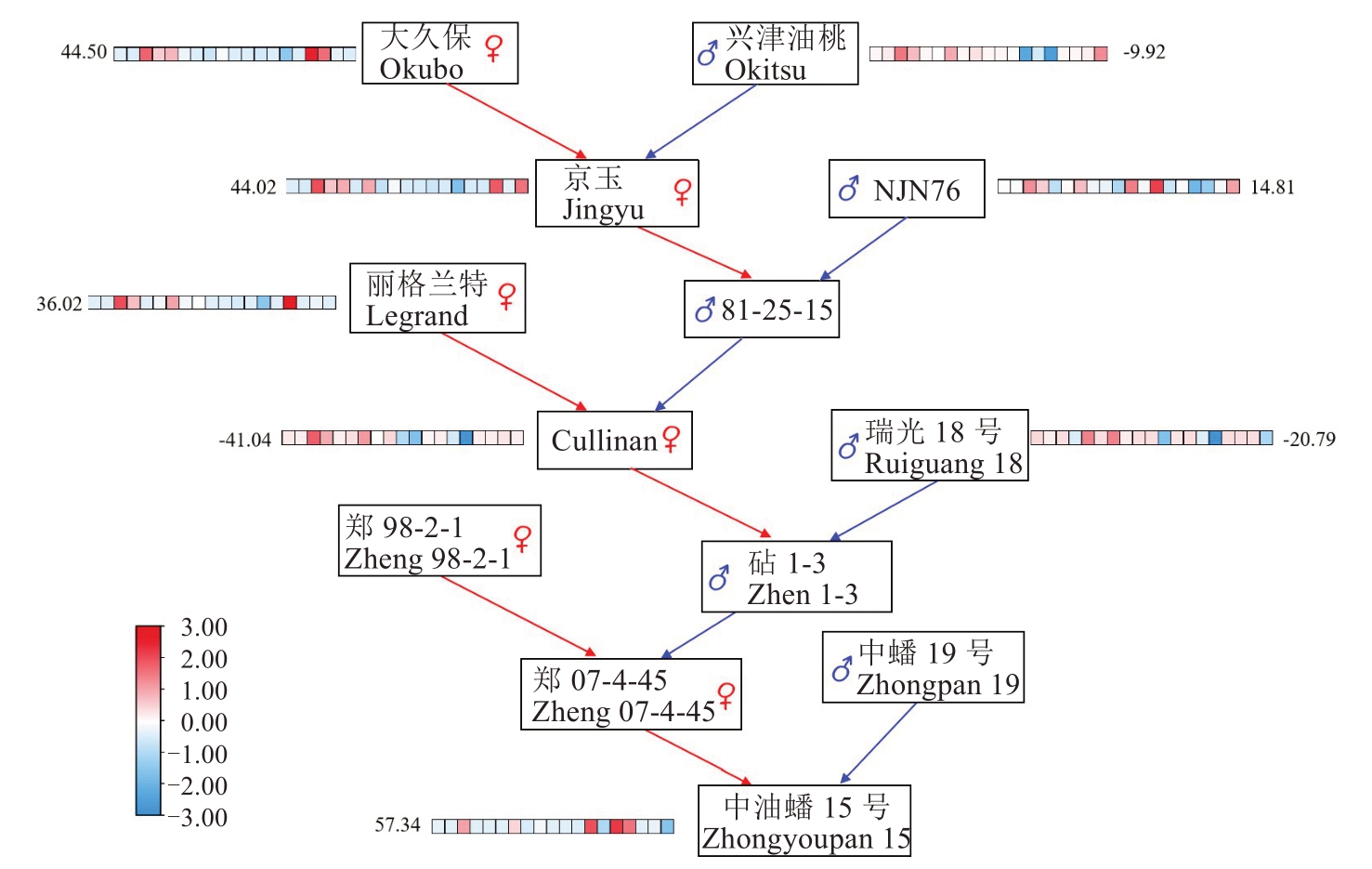
图3 中油蟠15号选育过程中亲本材料育种值的变化
Fig. 3 Changes in breeding values of parental materials during the breeding process of Zhong You Pan 15 peach
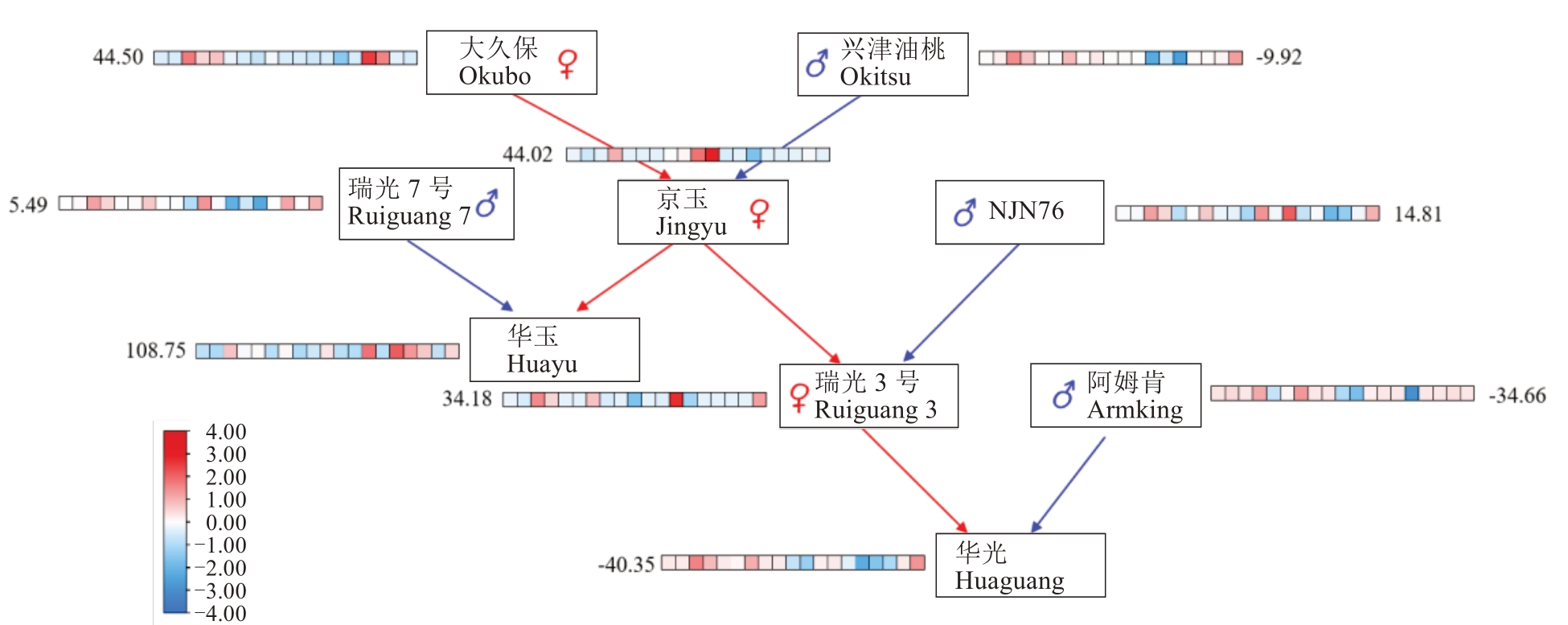
图4 华光选育过程中亲本材料育种值的变化
Fig. 4 Changes in breeding values of parental materials during the breeding process of Huaguang peach
2.4 主栽品种单果质量基因型组合的育种值与实测值的相关性分析
本研究选择了30个主栽品种(表3),计算出每个品种的预测育种值,并比较其与真实单果质量表型的相关性。结果表明,预测育种值与真实观测值之间呈极显著正相关,R2为0.467 3(r0.01,28=0.462 8,P=3.125 0E-05,图5)。应用19个位点来计算种质的育种值与其表型具有较好的相关性,可为单果质量性状的改良提供明确的分子标记选择支撑。
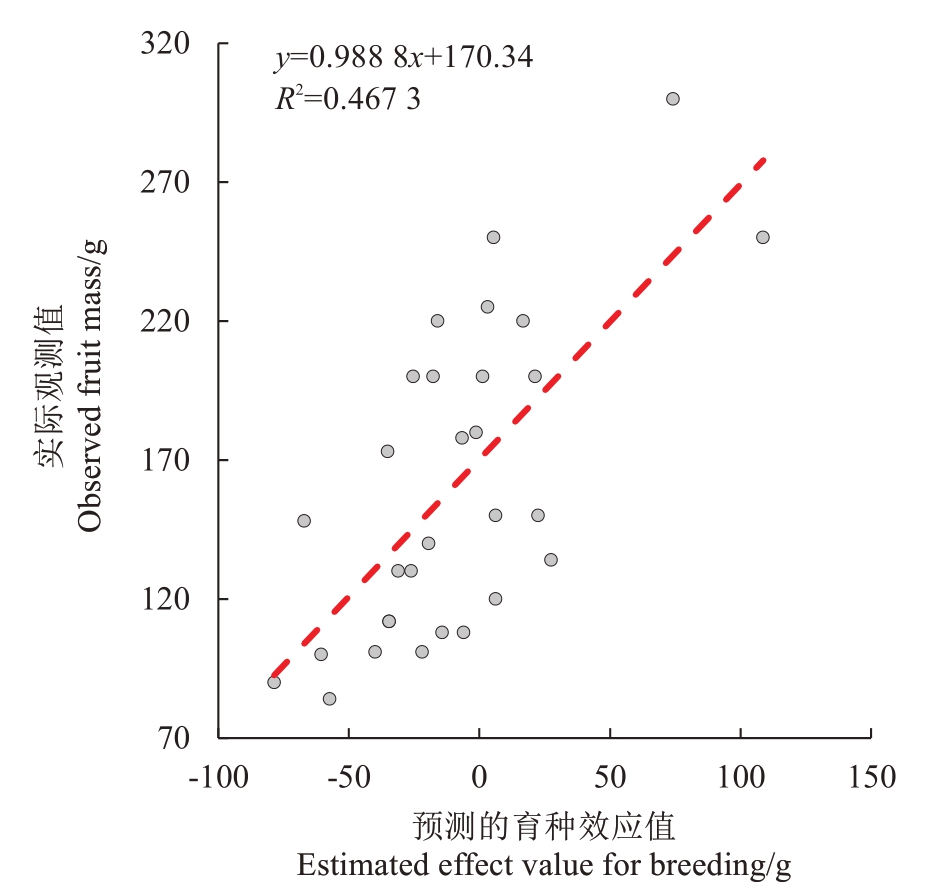
图5 预测育种值与实际观测的单果质量表型的相关性分析
Fig. 5 Correlation analysis between estimated breeding values and observed fruit mass
表3 30个主栽品种名称
Table 3 Thirty major cultivated variety names

?
2.5 筛选对主栽品种有改良意义的亲本种质
由于在30个主栽品种中,有24个品种均在效应值最高的N15位点为负效应,19个品种在效应值第2高的N13位点同样为负效应,因此,这些品种均可以与该位点为突变位点的种质(包含野生种质、地方品种或其他育种品种)进行杂交,通过基因型鉴定进行分子标记辅助筛选。在本研究中,育种值较高的前3个位点:在N15位点上,CC基因型的育种值达20.06,显著高于GG基因型,是提升单果质量的核心有利基因型,但在574份种质中仅占8.80%;N13位点的AA基因型育种值为18.04,显著优于GG基因型,其在574份种质中的占比也仅为8.95%,与N15位点共同构成单果质量改良的关键标记位点;N16位点的CC基因型育种值为14.84,显著高于TT基因型,该基因型在574份种质中占比相对较高(35.26%),但仍存在进一步优化的潜力。上述结果不仅明确了3个位点在高单果质量育种过程中利用的优先级,也为结合地方品种资源开展靶向选择提供了具体目标。对于低频的有利基因型位点(N15和N13),可优先从遗传背景丰富的地方品种中筛选携带该基因型的材料(长岭早玉露、白花、吉林8903、龙华水蜜桃、晚蟠桃和接土白等在N15和N13为正效应)作为亲本,通过定向杂交可以快速提升其在育种群体中的频率;而N16、N11、N3和N17等位点则可作为辅助选择标记,与前2个位点协同应用以实现单果质量性状的高效改良。
3 讨 论
桃单果质量是育种家关注的重要性状之一。本研究首先查阅并确定了国内外报道的46个连锁/关联标记,通过过滤筛选出19个独立的标记,利用这些标记计算单个位点的效应值进行叠加,也计算了3个不同品种杂交选育过程中亲本材料的育种值,以及30个主栽品种的育种值,表明育种值与实际观测的单果质量表型相关,证实这些标记可用于单果质量的分子辅助育种。然而,在华光的育种中,当以大果为育种目标时,其后代常因非加性效应(如显性/上位性)消失而出现果实变小的回归现象,这一规律已被多项研究证实[44-46]。此外,在番茄上,虽然Slclv3Pro-Slwuslc-Slcle9之间存在顺式调控层面的基因上位性关系,但联合效应往往不等于各基因独立效应的简单相加[47]。在水稻上,对18 421个水稻纯合品系(18K群体)研究显示,约34%的数量性状位点(QTLs)表现出显著的上位性效应,这些互作可解释16个核心农艺性状(千粒质量、单株产量、株高、分蘖角度、穗长等)15%~28%的表型变异[48]。在效应计算过程中,由于目前对位点间上位性效应与显性效应的解析仍存在技术与理论层面的局限,如相关效应的作用模式、强度及调控网络尚未完全阐明。因此,在分析现有单果质量的基因型组合总体效应值时,由于仅按照加性效应计算,会因忽略上位性效应和显性效应而产生一定偏差。在574份种质中,湖景蜜露、中华寿桃、中油蜜玉和中蟠17号等种质表现出大果表型,但基于现有加性效应计算得出的效应值却相对偏低。此外,也可能存在着其他的原因,如在水稻上,研究者从5000多份种质中鉴定到千粒质量超60 g的超大粒种质资源ULG,其除了包含了4个已报道的优势等位基因外,也发现了一个新的籽粒调节基因qULG2-b[49],即表型与育种值间的不匹配,与该种质存在不同于已知基因的新基因有关,这一现象在玉米[50]和葡萄[51]等研究中也有报道。
在基于上述分子标记开展亲本筛选以改良现有品种的过程中还需要考虑其他因素。龙华水蜜桃在单果质量相关表型及基因型检测中均呈现优良特征,符合高产育种的需求;然而,其携带的酸含量调控等位基因为劣势基因型,导致其在综合品质评价中表现欠佳。此外,某个优势基因型有可能与其他的劣势基因型存在连锁关系,进一步加大了育种难度。Xiao等[52]等通过水稻全基因组连锁分析,发现第6染色体Piz基因座的Hap1类型(感病等位基因)是制约产量、品质与抗病性协同改良的关键位点,亲本选择中若忽视该位点信息,将难以打破基因连锁壁垒,阻碍多性状聚合。在菜豆[53]上同样存在产量、花期及抗病性等农艺性状受多遗传位点协同调控的现象。因此,亲本筛选需通过全基因组重测序系统解析全基因组信息,通过全基因组关联分析鉴定各基因型对性状的遗传效应,并分析不同性状不同等位基因间的调控关系,为育种家提供全面准确的信息,推进综合性状优良的品种培育工作。
田间表型是由基因型、环境以及基因型与环境相互作用的结果。利用基因型精准鉴定来进行亲本选择时同样需要考虑环境因素。有研究提出,在构建基因组预测模型时,可分别构建代表遗传和环境因素的子模块,通过整合训练提升预测精度[54]。姚州等[55]开发的GEFormer方法,通过融合基因型与环境特征,在多个物种的多类群体中,预测准确率优于现有基因组预测方法。因此,若能在育种过程中有效利用环境因素,并通过开展多环境试验构建环境与基因型互作的模型,将显著提升育种效率,培育出适应不同环境、性状更为优良的桃品种。
4 结 论
桃单果质量是多基因控制的数量性状。本研究从已报道的46个连锁/关联标记中筛选得到19个独立标记,结合574份种质的基因组重测序数据,计算单个位点的效应值。利用这些标记计算3个不同品种杂交选育过程中亲本材料的育种值及30个主栽品种的育种值,结果显示育种值与实际观测的单果质量表型存在相关性。筛选出长岭早玉露、白花、吉林8903、晚蟠桃和接土白等对主栽品种有改良意义的亲本种质,为桃单果质量性状的育种工作提供了有价值的分子标记资源和技术支持。
[1] 曹云. 小麦春化发育基因型鉴定与相关基因的功能分析[D] .郑州:河南农业大学,2021.CAO Yun. Genotype identification and functional analysis of related genes in vernalization development of wheat[D] . Zhengzhou:Henan Agricultural University,2021.
[2] 王菲菲,张胜忠,胡晓辉,苗华荣,许林英,吴兰荣,李春娟,陈静. 16份花生品种(系)油酸含量及FAD2基因型鉴定[J] . 花生学报,2022,51(4):29-34.WANG Feifei,ZHANG Shengzhong,HU Xiaohui,MIAO Huarong,XU Linying,WU Lanrong,LI Chunjuan,CHEN Jing. The oleic content and FAD2 genotype identification of 16 peanut cultivars (lines)[J] . Journal of Peanut Science,2022,51(4):29-34.
[3] 赵雅楠,郑乾明,张敏,马玉华,赵德刚. 20个中国李品种自交不亲和S-RNɑse基因型的鉴定[J] . 中国果树,2023(11):37-42.ZHAO Yanan,ZHENG Qianming,ZHANG Min,MA Yuhua,ZHAO Degang. Identification of self-incompatibility S-RNɑse genotypes of 20 Chinese plum varieties[J] . China Fruits,2023(11):37-42.
[4] 刘春生,肖遥,范文琪,王宝安,徐亚,范嵩博,伍玲,鲜林峰,李威. 60个苹果新品种S基因型鉴定[J] . 河北果树,2023(1):5-7.LIU Chunsheng,XIAO Yao,FAN Wenqi,WANG Baoan,XU Ya,FAN Songbo,WU Ling,XIAN Linfeng,LI Wei. Identification of S-genotypes of 60 apple cultivars[J] . Hebei Fruits,2023(1):5-7.
[5] 张号楠,李沛华,梁东,王进,吕秀兰. 四川地区甜樱桃S基因型及SFB4’基因的分子鉴定[J] . 福建农业学报,2024,39(4):418-426.ZHANG Haonan,LI Peihua,LIANG Dong,WANG Jin,LÜ Xiulan. Identification of S genes and SFB4' in Cerɑsus ɑvium L.[J] .Fujian Journal of Agricultural Sciences,2024,39(4):418-426.
[6] 王朝晖,李勇,曹珂,朱更瑞,方伟超,陈昌文,王新卫,吴金龙,王力荣. 189份桃种质肉质性状形成相关位点基因型鉴定及组合分析[J] . 中国农业科学,2023,56(12):2367-2379.WANG Zhaohui,LI Yong,CAO Ke,ZHU Gengrui,FANG Weichao,CHEN Changwen,WANG Xinwei,WU Jinlong,WANG Lirong. Genotype identification and combination analysis of loci related to the peach flesh texture trait viɑ 189 peach accessions[J] . Scientia Agricultura Sinica,2023,56(12):2367-2379.
[7] 吉爽秋,王力荣,李勇,朱更瑞,曹珂,方伟超,陈昌文,王新卫,张琦,吴金龙. 桃花花型(铃形/蔷薇形)基因型鉴定、分子标记开发与利用[J] . 果树学报,2023,40(3):422-431.JI Shuangqiu,WANG Lirong,LI Yong,ZHU Gengrui,CAO Ke,FANG Weichao,CHEN Changwen,WANG Xinwei,ZHANG Qi,WU Jinlong. Identification of peach flower genotype (Nonshowy/Showy),development of flower-type related molecular markers[J] . Journal of Fruit Science,2023,40(3):422-431.
[8] 肖鑫,傅伟红,葛孟清,李腾,任艳华,马小河,刘崇怀,贾海锋,房经贵. 72个酿酒葡萄品种MYB调控基因的基因型和单倍型鉴定[J] . 南京农业大学学报,2022,45(1):27-36.XIAO Xin,FU Weihong,GE Mengqing,LI Teng,REN Yanhua,MA Xiaohe,LIU Chonghuai,JIA Haifeng,FANG Jinggui. Identification of genotypes and haplotypes of MYB-regulated genes in 72 wine grape varieties[J] . Journal of Nanjing Agricultural University,2022,45(1):27-36.
[9] FALCHI R,VENDRAMIN E,ZANON L,SCALABRIN S,CIPRIANI G,VERDE I,VIZZOTTO G,MORGANTE M.Three distinct mutational mechanisms acting on a single gene underpin the origin of yellow flesh in peach[J] . The Plant Journal,2013,76(2):175-187.
[10] DARDICK C,CALLAHAN A,HORN R,RUIZ K B,ZHEBENTYAYEVA T,HOLLENDER C,WHITAKER M,ABBOTT A,SCORZA R. PpeTAC1 promotes the horizontal growth of branches in peach trees and is a member of a functionally conserved gene family found in diverse plants species[J] .The Plant Journal,2013,75(4):618-630.
[11] VENDRAMIN E,PEA G,DONDINI L,PACHECO I,DETTORI M T,GAZZA L,SCALABRIN S,STROZZI F,TARTARINI S,BASSI D,VERDE I,ROSSINI L. A unique mutation in a MYB gene cosegregates with the nectarine phenotype in peach[J] .PLoS One,2014,9(3):e90574.
[12] PAN L,ZENG W F,NIU L,LU Z H,LIU H,CUI G C,ZHU Y Q,CHU J F,LI W P,FANG W C,CAI Z G,LI G H,WANG Z Q. PpYUC11,a strong candidate gene for the stony hard phenotype in peach (Prunus persicɑ L. Batsch),participates in IAA biosynthesis during fruit ripening[J] . Journal of Experimental Botany,2015,66(22):7031-7044.
[13] GU C,WANG L,WANG W,ZHOU H,MA B Q,ZHENG H Y,FANG T,OGUTU C,VIMOLMANGKANG S,HAN Y P. Copy number variation of a gene cluster encoding endopolygalacturonase mediates flesh texture and stone adhesion in peach[J] . Journal of Experimental Botany,2016,67(6):1993-2005.
[14] HOLLENDER C A,HADIARTO T,SRINIVASAN C,SCORZA R,DARDICK C. A brachytic dwarfism trait (dw) in peach trees is caused by a nonsense mutation within the gibberellic acid receptor PpeGID1c[J] . New Phytologist,2016,210(1):227-239.
[15] HOLLENDER C A,PASCAL T,TABB A,HADIARTO T,SRINIVASAN C,WANG W P,LIU Z C,SCORZA R,DARDICK C.Loss of a highly conserved sterile alpha motif domain gene(WEEP) results in pendulous branch growth in peach trees[J] .Proceedings of the National Academy of Sciences of the United States of America,2018,115(20):E4690-E4699.
[16] GUO J,CAO K,DENG C,LI Y,ZHU G R,FANG W C,CHEN C W,WANG X W,WU J L,GUAN L P,WU S,GUO W W,YAO J L,FEI Z J,WANG L R. An integrated peach genome structural variation map uncovers genes associated with fruit traits[J] . Genome Biology,2020,21(1):258.
[17] LU Z H,CAO H H,PAN L,NIU L,WEI B,CUI G C,WANG L W,YAO J L,ZENG W F,WANG Z Q. Two loss-of-function alleles of the glutathione S-transferase (GST) gene cause anthocyanin deficiency in flower and fruit skin of peach (Prunus persicɑ)[J] . The Plant Journal,2021,107(5):1320-1331.
[18] ZANON L,FALCHI R,SANTI S,VIZZOTTO G. Sucrose transport and phloem unloading in peach fruit:Potential role of two transporters localized in different cell types[J] . Physiologia Plantarum,2015,154(2):179-193.
[19] VIMOLMANGKANG S,ZHENG H Y,PENG Q,JIANG Q,WANG H L,FANG T,LIAO L,WANG L,HE H P,HAN Y P.Assessment of sugar components and genes involved in the regulation of sucrose accumulation in peach fruit[J] . Journal of Agricultural and Food Chemistry,2016,64(35):6723-6729.
[20] ASLAM M M,DENG L,WANG X B,WANG Y,PAN L,LIU H,NIU L,LU Z H,CUI G C,ZENG W F,WANG Z Q. Expression patterns of genes involved in sugar metabolism and accumulation during peach fruit development and ripening[J] . Scientia Horticulturae,2019,257:108633.
[21] WANG Q,CAO K,CHENG L L,LI Y,GUO J,YANG X W,WANG J,KHAN I A,ZHU G R,FANG W C,CHEN C W,WANG X W,WU J L,XU Q,WANG L R. Multi-omics approaches identify a key gene,PpTST1,for organic acid accumulation in peach[J] . Horticulture Research,2022,9:uhac026.
[22] ZHOU H,WANG K L,WANG H L,GU C,DARE A P,ESPLEY R V,HE H P,ALLAN A C,HAN Y P. Molecular genetics of blood-fleshed peach reveals activation of anthocyanin biosynthesis by NAC transcription factors[J] . The Plant Journal,2015,82(1):105-121.
[23] PAN L,LU Z H,YAN L L,ZENG W F,SHEN Z J,YU M L,BU L L,CUI G C,NIU L,WANG Z Q. NLR1 is a strong candidate for the Rm3 dominant green peach aphid (Myzus persicɑe)resistance trait in peach[J] . Journal of Experimental Botany,2022,73(5):1357-1369.
[24] CAO K,PENG Z,ZHAO X,LI Y,LIU K Z,ARUS P,FANG W C,CHEN C W,WANG X W,WU J L,FEI Z J,WANG L R.Chromosome-level genome assemblies of four wild peach species provide insights into genome evolution and genetic basis of stress resistance[J] . BMC Biology,2022,20(1):139.
[25] DIRLEWANGER E,MOING A,ROTHAN C,SVANELLA L,PRONIER V,GUYE A,PLOMION C,MONET R. Mapping QTLs controlling fruit quality in peach [Prunus persicɑ (L.)Batsch] [J] . Theoretical and Applied Genetics,1999,98(1):18-31.
[26] ETIENNE C,ROTHAN C,MOING A,PLOMION C,BODÉNÈS C,SVANELLA-DUMAS L,COSSON P,PRONIER V,MONET R,DIRLEWANGER E. Candidate genes and QTLs for sugar and organic acid content in peach [Prunus persicɑ (L.) Batsch] [J] . Theoretical and Applied Genetics,2002,105(1):145-159.
[27] QUILOT B,WU B H,KERVELLA J,GÉNARD M,FOULONGNE M,MOREAU K. QTL analysis of quality traits in an advanced backcross between Prunus persicɑ cultivars and the wild relative species P. dɑvidiɑnɑ[J] . Theoretical and Applied Genetics,2004,109(4):884-897.
[28] EDUARDO I,PACHECO I,CHIETERA G,BASSI D,POZZI C,VECCHIETTI A,ROSSINI L. QTL analysis of fruit quality traits in two peach intraspecific populations and importance of maturity date pleiotropic effect[J] . Tree Genetics & Genomes,2011,7(2):323-335.
[29] DA SILVA LINGE C,BASSI D,BIANCO L,PACHECO I,PIRONA R,ROSSINI L. Genetic dissection of fruit weight and size in an F2 peach [Prunus persicɑ (L.) Batsch] progeny[J] . Molecular Breeding,2015,35(2):71.
[30] ZEBALLOS J L,ABIDI W,GIMÉNEZ R,MONFORTE A J,MORENO M Á,GOGORCENA Y. Mapping QTLs associated with fruit quality traits in peach [Prunus persicɑ (L.) Batsch] using SNP maps[J] . Tree Genetics & Genomes,2016,12(3):37.
[31] SHI P,XU Z,ZHANG S Y,WANG X J,MA X F,ZHENG J C,XING L B,ZHANG D,MA J J,HAN M Y,ZHAO C P. Construction of a high-density SNP-based genetic map and identification of fruit-related QTLs and candidate genes in peach[Prunus persicɑ (L.) Batsch] [J] . BMC Plant Biology,2020,20(1):438.
[32] CAO K,ZHOU Z K,WANG Q,GUO J,ZHAO P,ZHU G R,FANG W C,CHEN C W,WANG X W,WANG X L,TIAN Z X,WANG L R. Genome-wide association study of 12 agronomic traits in peach[J] . Nature Communications,2016,7:13246.
[33] CAO K,LI Y,DENG C H,GARDINER S E,ZHU G R,FANG W C,CHEN C W,WANG X W,WANG L R. Comparative population genomics identified genomic regions and candidate genes associated with fruit domestication traits in peach[J] .Plant Biotechnology Journal,2019,17(10):1954-1970.
[34] KSOURI N,SANCHEZ G,FORCADA C F I,CONTRERASMOREIRA B,GOGORCENA Y. ddRAD-seq-derived SNPs reveal novel association signatures for fruit-related traits in peach[J] .bioRxiv,2023. https://doi.org/10.1101/2023.07.31.551252.
[35] WANG J T,CHEN Y,SHU G P,ZHAO M M,ZHENG A,CHANG X Y,LI G Q,WANG Y B,ZHANG Y M. Fast3VmrMLM:A fast algorithm that integrates genome-wide scanning with machine learning to accelerate gene mining and breeding by design for polygenic traits in large-scale GWAS datasets[J] .Plant Communications,2025,6(7):101385.
[36] ZHANG Y Y,WANG Y J,HENKE M,CARBONELL-BEJERANO P,WANG Z M,BERT P F,WANG Y,LI H Y,KONG J H,FAN P G,DAI Z W,LIANG Z C. Integrating dense genotyping with high-throughput phenotyping empowers the genetic dissection of berry quality and resilience traits in grapevine[J] . Advanced Science,2025,12(29):2412587.
[37] WANG F M,LIU Y,ZHANG A N,KONG D Y,BI J G,LIU G L,YU X Q,LUO L J. Breeding an early maturing,blast resistance water-saving and drought-resistance rice (WDR) cultivar using marker-assisted selection coupled with rapid generation advance[J] . Molecular Breeding,2022,42(8):46.
[38] 王力荣,朱更瑞,方伟超. 中国桃遗传资源[M] . 北京:中国农业出版社,2012.WANG Lirong,ZHU Gengrui,FANG Weichao. Peach genetic resource in China[M] . Beijing:China Agriculture Press,2012.
[39] BOLGER A M,LOHSE M,USADEL B. Trimmomatic:A flexible trimmer for Illumina sequence data[J] . Bioinformatics,2014,30(15):2114-2120.
[40] LI H. Aligning sequence reads,clone sequences and assembly contigs with BWA-MEM[J] . ArXiv:Genomics,2013. https://doi.org/10.48550/arXiv.1303.3997.
[41] MCKENNA A,HANNA M,BANKS E,SIVACHENKO A,CIBULSKIS K,KERNYTSKY A,GARIMELLA K,ALTSHULER D,GABRIEL S,DALY M,DEPRISTO M A. The Genome Analysis Toolkit:A MapReduce framework for analyzing nextgeneration DNA sequencing data[J] . Genome Research,2010,20(9):1297-1303.
[42] LI Y,CAO K,ZHU G R,FANG W C,CHEN C W,WANG X W,ZHAO P,GUO J,DING T Y,GUAN L P,ZHANG Q,GUO W W,FEI Z J,WANG L R. Genomic analyses of an extensive collection of wild and cultivated accessions provide new insights into peach breeding history[J] . Genome Biology,2019,20(1):36.
[43] DA SILVA LINGE C,CAI L C,FU W F,CLARK J,WORTHINGTON M,RAWANDOOZI Z,BYRNE D H,GASIC K. Multi-locus genome-wide association studies reveal fruit quality hotspots in peach genome[J] . Frontiers in Plant Science,2021,12:644799.
[44] 韩明玉. 桃主要性状遗传分析与再生体系的建立[D] . 杨凌:西北农林科技大学,2009.HAN Mingyu. Genetic analysis of main traits and establishment of regeneration system of peach[D] . Yangling:Northwest A & F University,2009.
[45] 王逢寿,刘桂林. 桃、樱桃主要经济性状的遗传[J] . 辽宁果树,1979(1):18-22.WANG Fengshou,LIU Guilin. Inheritance of main economic characters of peach and cherry[J] . Liaoning Fruits,1979(1):18-22.
[46] 浙江农业大学. 果树育种学[M] . 上海:上海科学技术出版社,1980.Zhejiang Agricultural University. Fruit tree breeding[M] . Shanghai:Shanghai Scientific & Technical Publishers,1980.
[47] AGUIRRE L,HENDELMAN A,HUTTON S F,MCCANDLISH D M,LIPPMAN Z B. Idiosyncratic and dose-dependent epistasis drives variation in tomato fruit size[J] . Science,2023,382(6668):315-320.
[48] WEI X,CHEN M J,ZHANG Q,GONG J Y,LIU J,YONG K C,WANG Q,FAN J J,CHEN S H,HUA H,LUO Z W,ZHAO X Y,WANG X,LI W,CONG J,YU X T,WANG Z H,HUANG R P,CHEN J X,ZHOU X Y,QIU J,XU P,MURRAY J,WANG H,XU Y,XU C W,XU G,YANG J L,HAN B,HUANG X H. Genomic investigation of 18,421 lines reveals the genetic architecture of rice[J] . Science,2024,385(6704):eadm8762.
[49] LI F C,WU L,LI X,CHAI Y N,RUAN N,WANG Y,XU N,YU Z W,WANG X C,CHEN H,LU J H,XU H,XU Z J,CHEN W F,XU Q. Dissecting the molecular basis of the ultralarge grain formation in rice[J] . New Phytologist,2024,243(6):2251-2264.
[50] WANG B B,HOU M,SHI J P,KU L X,SONG W,LI C H,NING Q,LI X,LI C Y,ZHAO B B,ZHANG R Y,XU H,BAI Z J,XIA Z C,WANG H,KONG D X,WEI H B,JING Y F,DAI Z Y,WANG H H,ZHU X Y,LI C H,SUN X,WANG S S,YAO W,HOU G G,QI Z,DAI H,LI X M,ZHENG H K,ZHANG Z X,LI Y,WANG T Y,JIANG T J,WAN Z M,CHEN Y H,ZHAO J R,LAI J S,WANG H Y. De novo genome assembly and analyses of 12 founder inbred lines provide insights into maize heterosis[J] . Nature Genetics,2023,55(2):312-323.
[51] LIU Z J,WANG N,SU Y,LONG Q M,PENG Y L,SHANGGUAN L F,ZHANG F,CAO S,WANG X,GE M Q,XUE H,MA Z Y,LIU W W,XU X D,LI C C,CAO X J,AHMAD B,SU X N,LIU Y T,HUANG G Z,DU M R,LIU Z Y,GAN Y,SUN L,FAN X C,ZHANG C,ZHONG H X,LENG X P,REN Y H,DONG T Y,PEI D,WU X Y,JIN Z X,WANG Y W,LIU C H,CHEN J F,GAUT B,HUANG S W,FANG J G,XIAO H,ZHOU Y F. Grapevine pangenome facilitates trait genetics and genomic breeding[J] . Nature Genetics,2024,56(12):2804-2814.
[52] XIAO N,PAN C H,LI Y H,WU Y Y,CAI Y,LU Y,WANG R Y,YU L,SHI W,KANG H X,ZHU Z B,HUANG N S,ZHANG X X,CHEN Z C,LIU J J,YANG Z F,NING Y S,LI A H. Genomic insight into balancing high yield,good quality,and blast resistance of Japonica rice[J] . Genome Biology,2021,22(1):283.
[53] WU J,WANG L F,FU J J,CHEN J B,WEI S H,ZHANG S L,ZHANG J,TANG Y S,CHEN M L,ZHU J F,LEI L,GENG Q H,LIU C L,WU L,LI X M,WANG X L,WANG Q,WANG Z L,XING S L,ZHANG H K,BLAIR M W,WANG S M. Resequencing of 683 common bean genotypes identifies yield component trait associations across a north-south cline[J] . Nature Genetics,2020,52(1):118-125.
[54] HE K H,YU T X,GAO S,CHEN S K,LI L,ZHANG X C,HUANG C L,XU Y B,WANG J K,PRASANNA B M,HEARNE S,LI X H,LI H H. Leveraging automated machine learning for environmental data-driven genetic analysis and genomic prediction in maize hybrids[J] . Advanced Science,2025,12(17):2412423.
[55] YAO Z,YAO M T,WANG C,LI K,GUO J H,XIAO Y J,YAN J B,LIU J X. GEFormer:A genotype-environment interactionbased genomic prediction method that integrates the gating multilayer perceptron and linear attention mechanisms[J] . Molecular Plant,2025,18(3):527-549.