桃[Prunus persica (L.) Batsch]是深受人们喜爱的世界性大宗果品,在全球南、北纬30°~45°的广大范围内均有栽培,果实品质的优劣直接影响其经济价值。生长素在桃果实的发育过程中扮演着重要角色。前期研究表明,在桃果实缝合线软化过程中,内源激素含量发生显著变化,其中生长素被证实对中果皮细胞的膨大及成熟软化过程具有重要调控作用[1]。对油桃24-30 的采后生长素处理发现,果实的乙烯释放量迅速增加,果实硬度明显下降,且乙烯生物合成及果实软化途径中关键基因的表达量显著增加,这表明生长素可以通过调控相关基因的表达促进乙烯释放,加速果实软化进程[2]。对不同品种桃果实中生长素含量变化的研究显示,裂核桃果实中IAA 含量明显高于正常果实,而果核的硬化和开裂与木质素的合成直接相关,这表明生长素可以通过影响木质素的合成来调控桃果核的发育,进而影响桃果实的品质[3]。
生长素信号转导是调控高等植物生长发育的核心机制[4],该过程起始于F-box 蛋白受体(TIR1/AFB)对生长素的感知,其与Skp1、Cullin 蛋白组装形成SCF型E3泛素连接酶复合体[5]。在生长素信号诱导下,该复合体催化AUX/IAA 蛋白的泛素化降解[5-6]。AUX/IAA 蛋白家族作为转录抑制因子,通过抑制生长素响应因子(ARFs)的活性,在生长素信号转导中发挥核心调控作用[4,7-8]。目前,拟南芥、草莓、苹果及桃等多种植物的AUX/IAA 基因家族已被系统鉴定[9-11],多项研究表明该家族成员在果实发育的不同阶段发挥重要功能[12-13]。已有研究在桃基因组中鉴定出23 个AUX/IAA 基因[2,14]。在晚24 号桃果实发育中后期,PpIAA11 的表达水平随果实成熟进程而逐渐上升[15]。在番茄中,与PpIAA11 高度同源的SlIAA13 在外源生长素处理后表达显著上调[16-17],并可结合转录激活因子ARF5 和ARF6,负调控生长素信号转导过程[18]。此外,在番茄中异源过表达PpIAA19 能够影响植株形态及果实发育[12]。目前,对桃AUX/IAA 基因家族中单个基因功能的深入研究仍相对有限。
目前,我国鲜食桃市场以溶质型品种为主,然而果实采后软化迅速,严重制约了其贮藏与运输性能[2,15]。因此,深入研究桃果实发育与成熟的调控机制对产业实践具有重要意义。基于日川白凤桃不同发育时期的转录组数据发现,PpIAA11(Prupe.8G215400)在花后70 d(即第2 次快速膨大期)的表达量最高,表明该基因可能参与调控果实发育进程[15]。已有研究表明,外源施用萘乙酸(1-Naphthaleneacetic acid,NAA)及生长素转运抑制剂萘基邻苯二甲酸(N-1-napthylphthalamic acid,NPA)可通过影响乙烯合成进而调控桃果实的发育与成熟进程[19-21],且该调控效应具有处理时期和浓度依赖性[22-24]。例如,在果实成熟阶段,生长素促进乙烯合成的作用随处理浓度升高而增强[24],然而在花后30 d时,低浓度NAA(0.25 mmol·L-1)促进果实成熟,而较高浓度NAA(0.5、1、2 mmol·L-1)则抑制成熟进程,甚至导致果实畸形与脱落[20]。因此,为探究PpIAA11 对外源生长素类物质的响应模式及其在桃果实发育中的潜在功能,以花后70 d的桃果实为试材,采用不同浓度的NAA 与NPA 进行处理。通过克隆PpIAA11 基因,开展生物信息学分析、表达模式解析及番茄异源转化等试验,系统研究了PpIAA11 在响应生长素信号并参与调控桃果实发育过程中的作用。
1 材料和方法
1.1 试验材料及处理
以10 年生的日川白凤水蜜桃树为材料,分别于谢花后20 d(第1次快速生长期)、50 d(硬核期)、70 d(第2 次快速生长期)和90 d(成熟期)采集果实样品。基于前期的研究基础[24-25],选择在花后70 d采用浓度为0.1、1.0和2.0 mmol·L-1的NAA(含1% 吐温-80)及NPA(含1% 吐温-80)对果实进行处理[25-26],以等量1% 吐温-80 的清水作为对照。分别于处理后6、12、24、48 和72 h 动态采集果实样品,每个处理设置3 个生物学重复。所有样品经液氮速冻后置于-80 ℃超低温冰箱中保存备用。
1.2 桃果实总RNA提取及cDNA合成
总RNA 提取采用TaKaRa生物技术有限公司的通用型试剂盒,获得的总RNA 经质量检测合格后,使用PrimeScript™ RT reagent Kit with gDNA Eraser(Perfect Real Time)反转录试剂盒(TaKaRa)合成cDNA,产物于-20 ℃冰箱中保存备用。
1.3 PpIAA11基因克隆
基于团队前期的转录组测序结果,筛选得到PpIAA11 基因,并在桃基因组数据库(https://www.rosaceae.org/organism)中获取其cDNA 参考序列。依据该基因的CDS 区段,使用Primer Premier 5.0 软件设计特异性PCR 扩增引物(引物序列见表1),以合成的cDNA 为模板进行PCR 扩增[27]。反应程序如下:98 ℃变性10 s,56 ℃退火15 s,72 ℃延伸30 s,共35 个循环。PCR 产物经1%琼脂糖凝胶电泳分离,在凝胶成像系统中检测并确认目的条带,随后使用北京天根生化科技有限公司的DNA 凝胶回收试剂盒进行纯化。将纯化产物与pMD19-T 载体连接,并转化至大肠杆菌(Escherichia coli)DH5α 感受态细胞。经培养后,挑取单克隆进行菌落PCR 鉴定,并对阳性克隆进行测序验证。试验所用Taq 酶、dNTPs、pMD19-T 载体、DNA Marker 及大肠杆菌菌株DH5α均购自TaKaRa公司。
表1 试验中所用引物序列
Table 1 Primer sequence used in the experiment

引物名称Primer name PpIAA11 PpIAA11-qRT-PCR PpACTIN-qRT-PCR PpIAA11-35S上游引物Forward primer (5′-3′)ATGGAGGGTGTTTTGGGTAGTG GGGAAGGCAAACAGTGGAAA ATTCCCTGACTGTTTGCTAGT AGAACACGGGGGACGAGCTCATGGAGGGTGTTTTGGGTAGTGG下游引物Reverse primer (5′-3′)CAAAGATGCCAGCCGATATAG CCCTGACCACCTGAACGTAT TCCAACACAATACCGGTGGT ACCATGGTGTCGACTCTAGATATCGGCTGGCATCTTTGCCTCA
1.4 PpIAA11序列分析
利用PpIAA11 氨基酸序列,在NCBI BLAST 网站上搜索与其同源性最高的果树作物IAA11 序列。利用DNAMAN 软件进行多重序列比对,并采用MEGA 11.0 软件以邻接法(neighbor-joining method)构建系统发育树,bootstrap 值设为1000。通过MEME 在线程序(http://meme-suite.org/)对PpIAA11蛋白的保守基序进行分析。通过NCBI 保守结构域数据库(https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi)对该蛋白的保守结构域进行注释。使用PlantCARE(http://bioinformat cs.psb.ugent.be/webtools/plantCARE/html)和New PLACE 在线网站(https://www.dna.affrc.go.jp/PLACE/action=newplace)对PpIAA11 启动子序列进行顺式作用元件分析,并综合比对两个数据库的分析结果,以相互验证后的数据作为最终分析依据。
1.5 PpIAA11基因的表达分析
通过qRT-PCR 技术分析桃果实中PpIAA11的表达水平,所用定量引物及内参基因(Actin)引物序列见表1。每个试验设置3次重复,采用2-ΔΔCt方法计算基因的相对表达量。所有试验数据使用Excel 2010软件进行整理与分析。
1.6 PpIAA11基因过表达载体构建
将经测序验证正确的阳性质粒与pCAMBIA2300-GFP 空载体,利用SacI和XbaI进行双酶切处理。以pCAMBIA2300-GFP 载体为骨架设计引物,在其上下游引物中分别引入SacI和XbaI酶切位点接头序列(引物见表1)。使用南京诺唯赞生物科技股份有限公司的ClonExpress® Ⅱ One Step Cloning Kit(C112,Vazyme)试剂盒进行无缝克隆连接。将连接产物通过热激法转化至大肠杆菌DH5α 感受态细胞,涂板培养过夜。挑取单克隆进行菌落PCR鉴定,并对阳性克隆进行测序验证[28]。
1.7 番茄遗传转化、筛选及鉴定
通过根癌农杆菌GV3101 介导的叶盘法对Micro-Tom 番茄进行遗传转化,获得T0 代转基因幼苗。将T0代植株移栽至花盆中培养,采集叶片并提取基因组DNA,通过PCR 技术进行阳性植株鉴定。将收获T0 代自交种子作为T1 代,播种T1 代及野生型(WT)番茄种子。待幼苗生长至适宜时期,采集叶片并采用CTAB 法提取基因组DNA[12],对T1代植株进行PCR 阳性鉴定。最终,对鉴定为阳性的转基因植株及WT对照植株进行果实表型观察与比较分析。
2 结果与分析
2.1 PpIAA11基因克隆及其表达分析
以桃果肉cDNA 为模板,设计引物并进行PCR扩增,获得与预期大小一致的目的条带(图1-A)。测序结果表明,PpIAA11 的基因编码区(CDS)全长939 bp,编码312 个氨基酸。qRT-PCR 结果分析显示,该基因在桃果实不同发育时期的表达量存在显著差异(图1-B)。其中,在花后70 d 时(第2 次快速膨大期)表达量最高,该结果与团队前期转录组数据一致[15],表明PpIAA11可能在桃果实第2次快速膨大过程中发挥重要调控作用。
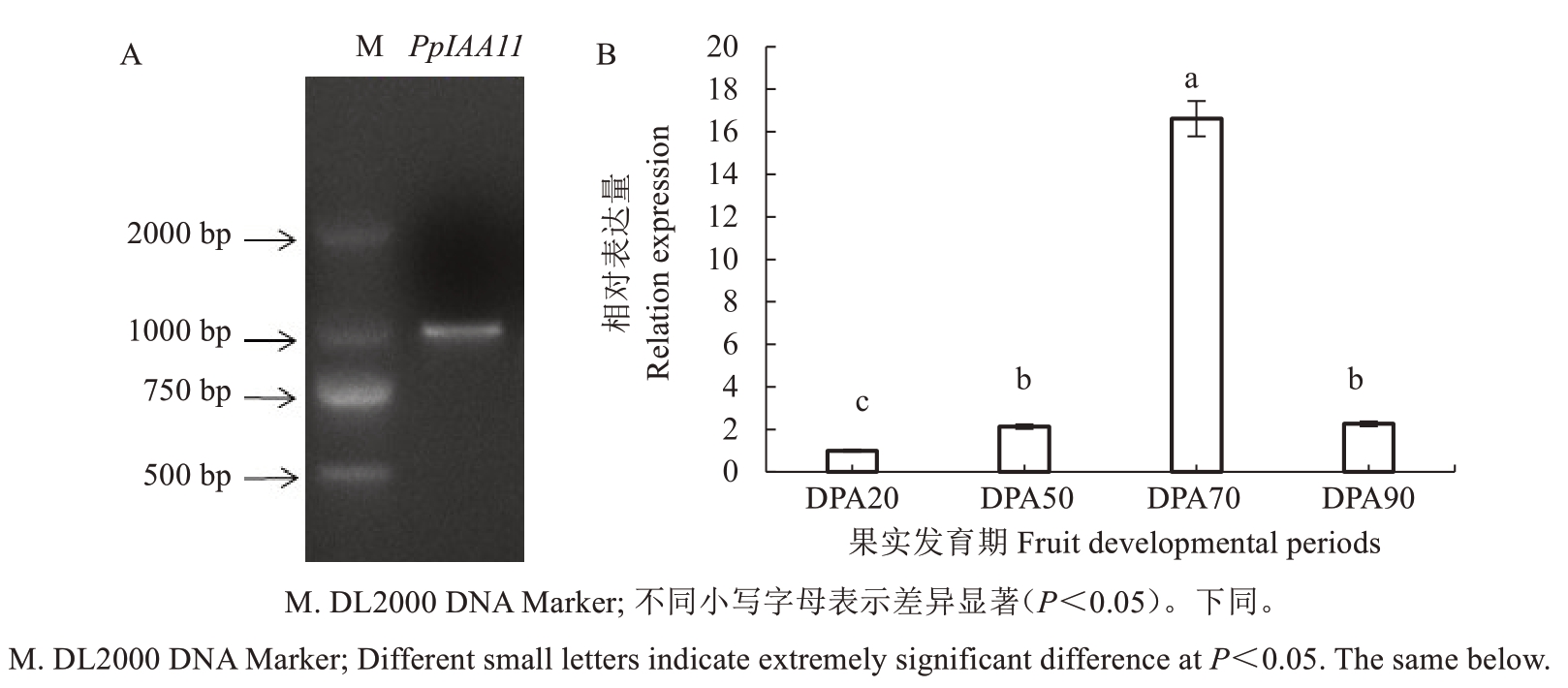
图1 PpIAA11 基因的PCR 扩增产物检测(A)及其在果实不同发育时期的表达(B)
Fig. 1 Detection of PCR amplified products of PpIAA11 gene (A) and its expression at different developmental periods of fruit (B)
2.2 PpIAA11蛋白序列分析
NCBI保守结构域分析显示,PpIAA11的氨基酸序列包含一个典型的AUX_IAA 结构域(登录号:pfam02309)。为探究其进化关系,将PpIAA11与9种果树作物的IAA11同源蛋白进行多重序列比对,包括巴旦木(Prunus dulcis,XP_034227148.1,PdIAA11)、欧洲甜樱桃(Prunus avium,XP_021831332.1,PaIAA11)、苹果(Malus domestica,NP_001315716.1,MdIAA11)、梨(Pyrus × bretschneideri,XP_009335227.2,PbIAA11)、河岸葡萄(Vitis riparia,XP_034692264.1,VrIAA11)、美国山核桃(Carya illinoinensis,XP_042974400.1,CiIAA11)、核桃(Juglans regia,XP_018807881.1,JrIAA11)、欧榛(Corylus avellana,XP_059432402.1,CaIAA11)和酸枣(Ziziphus jujuba var.spinosa,XP_015901174.1,ZjIAA11)。结果显示PpIAA11 与PdIAA11、PaIAA11 和MdIAA11 的序列相似性均高于95%,与其他6 个同源蛋白的相似性平均超过80%(图2-A)。其中,第111~299位氨基酸为高度保守区域,该区段与AUX_IAA 结构域范围基本一致,表明其为该家族功能核心区域。

图2 PpIAA11 与其他物种同源蛋白序列的多重比对(A)、AUX_IAA 结构域分析(B)及系统进化树构建(C)
Fig. 2 Multiple sequence alignment of PpIAA11 with homologous proteins from other species (A), AUX_IAA domain analysis (B), and construction of the phylogenetic tree (C)
为验证PpIAA11 的保守性,笔者进一步比较了不同物种中AUX_IAA 蛋白结构域(图2-B)。结果显示,与PpIAA11 类似,其余9 个果树物种的同源蛋白C 端附近均包含一个AUX_IAA 结构域,在不同物种中该结构域长度略有差异,PpIAA11、PdIAA11和PaIAA11 均由188 个氨基酸组成,CiIAA11 和JrIAA11 为191 个氨基酸,而MdIAA11、PbIAA11、CaIAA11 和ZjIAA11 分别为184、212、189 和192 个氨基酸。不同物种中AUX_IAA 结构域位置上的相似性,表明它是AUX_IAA 蛋白家族的一个显著特征。基于上述10 个物种AUX/IAA 蛋白序列构建的系统进化树显示,这些同源蛋白可分为两组(图2-C)。第一组仅包含VrIAA11,第二组则包含其余所有序列,并可进一步划分为两个亚组,亚组1 包括PpIAA11、PdIAA11、PaIAA11、MdIAA11 和PbIAA11;亚组2 包括ZjIAA、CaIAA11、CiIAA11 和JrIAA11。值得注意的是,PpIAA11 与PaIAA11 和PdIAA11 聚为一簇,表明它们的亲缘关系较近,该结果与序列相似性比对结论一致。
2.3 PpIAA11启动子顺式元件分析
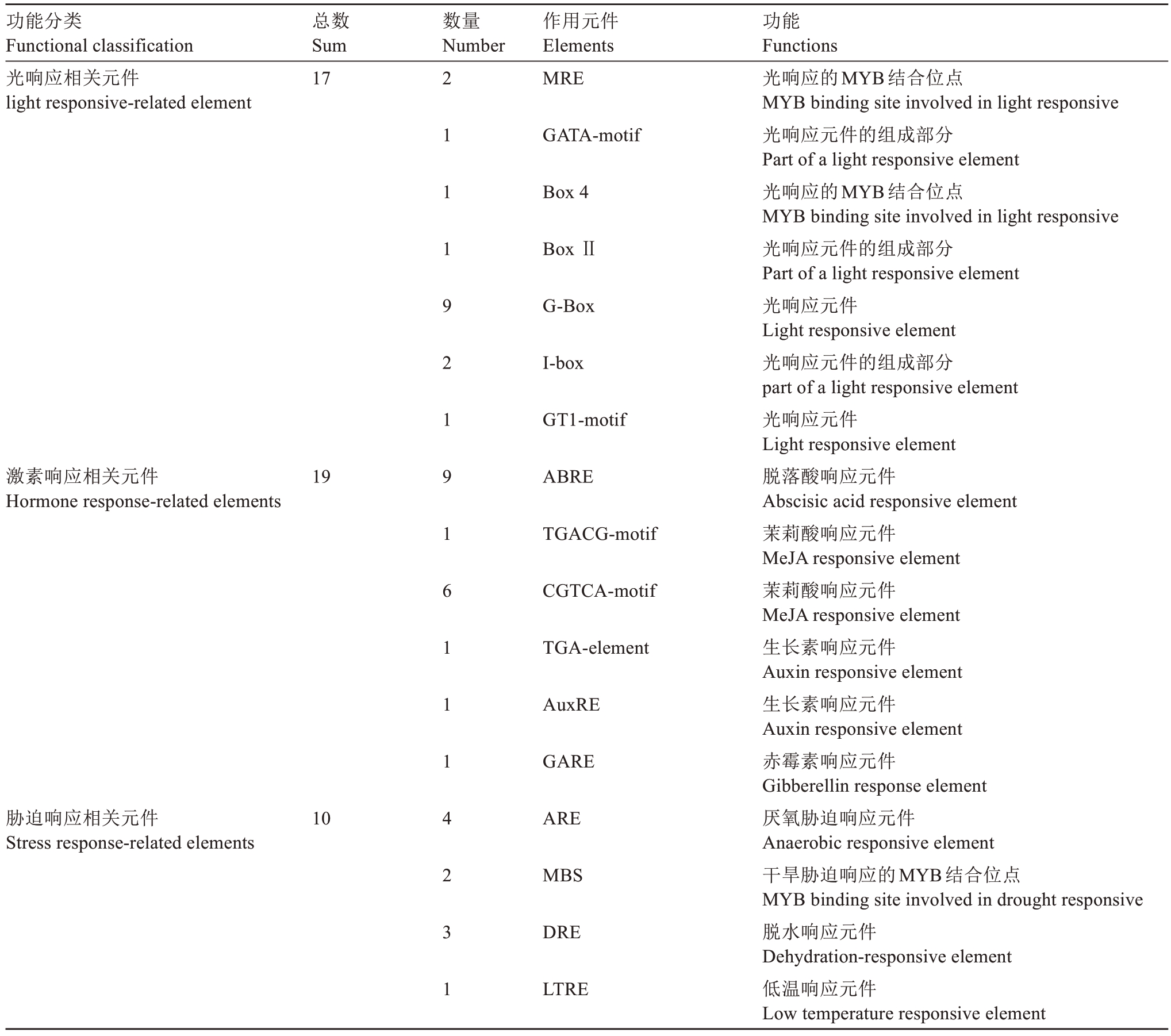
为预测PpIAA11 的潜在功能,利用PlantCARE和New PLACE 在线工具对其翻译起始位点上游2 kb的启动子区域进行了顺式作用元件分析。结果(表2)表明,PpIAA11 启动子区域主要包含光响应(17 个)、激素响应(19 个)及胁迫响应(10 个)元件。在激素响应元件中,包括9 个脱落酸(ABA)响应元件、7 个茉莉酸甲酯(MeJA)响应元件、2 个生长素(IAA)响应元件及1 个赤霉素(GA)响应元件。以上结果表明,PpIAA11 可能通过响应多种激素信号,在桃的生长发育过程中参与激素介导的转录调控。
表2 PpIAA11 基因启动子区域顺式作用元件分析
Table 2 The cis-acting elements analysis in the promoter regions of PpIAA11
功能分类Functional classification光响应相关元件light responsive-related element总数Sum 17数量Number作用元件Elements MRE GATA-motif Box 4 Box ⅡG-Box I-box GT1-motif激素响应相关元件Hormone response-related elements 19 ABRE TGACG-motif CGTCA-motif TGA-element AuxRE GARE胁迫响应相关元件Stress response-related elements 10 2 1 1 1 9 2 1 9 1 6 1 1 1 4 2 3 1 ARE MBS DRE LTRE功能Functions光响应的MYB结合位点MYB binding site involved in light responsive光响应元件的组成部分Part of a light responsive element光响应的MYB结合位点MYB binding site involved in light responsive光响应元件的组成部分Part of a light responsive element光响应元件Light responsive element光响应元件的组成部分part of a light responsive element光响应元件Light responsive element脱落酸响应元件Abscisic acid responsive element茉莉酸响应元件MeJA responsive element茉莉酸响应元件MeJA responsive element生长素响应元件Auxin responsive element生长素响应元件Auxin responsive element赤霉素响应元件Gibberellin response element厌氧胁迫响应元件Anaerobic responsive element干旱胁迫响应的MYB结合位点MYB binding site involved in drought responsive脱水响应元件Dehydration-responsive element低温响应元件Low temperature responsive element
2.4 PpIAA11对生长素处理的响应表达分析
为探究PpIAA11 对生长素信号的响应模式,分析了经不同浓度(0.1、1.0、2.0 mmol·L-1)NAA 和NPA处理后,PpIAA11在不同时间(6、12、24、48、72 h)的表达量动态变化。结果显示,0.1 mmol·L-1 NAA处理可诱导PpIAA11 的表达量在6 h 时短暂上升,随后急剧下降,至24 h 时降至最低,并持续显著低于对照组。1.0 mmol·L-1 NAA 处理6 h 和12 h 时,PpIAA11 表达量上调,之后逐渐回落至低水平。而2.0 mmol·L-1 NAA 处理则在整个时间范围内显著抑制PpIAA11 表达,但未表现出明显的时间依赖性变化。在NPA 处理组中,0.1 mmol·L-1 NPA 即可引起PpIAA11 的表达量显著下调,并在24 h 时达到最低;1.0 mmol·L-1和2.0 mmol·L-1 NPA 处理也持续抑制其表达,但不同时间点间变化趋势不明显(图3)。
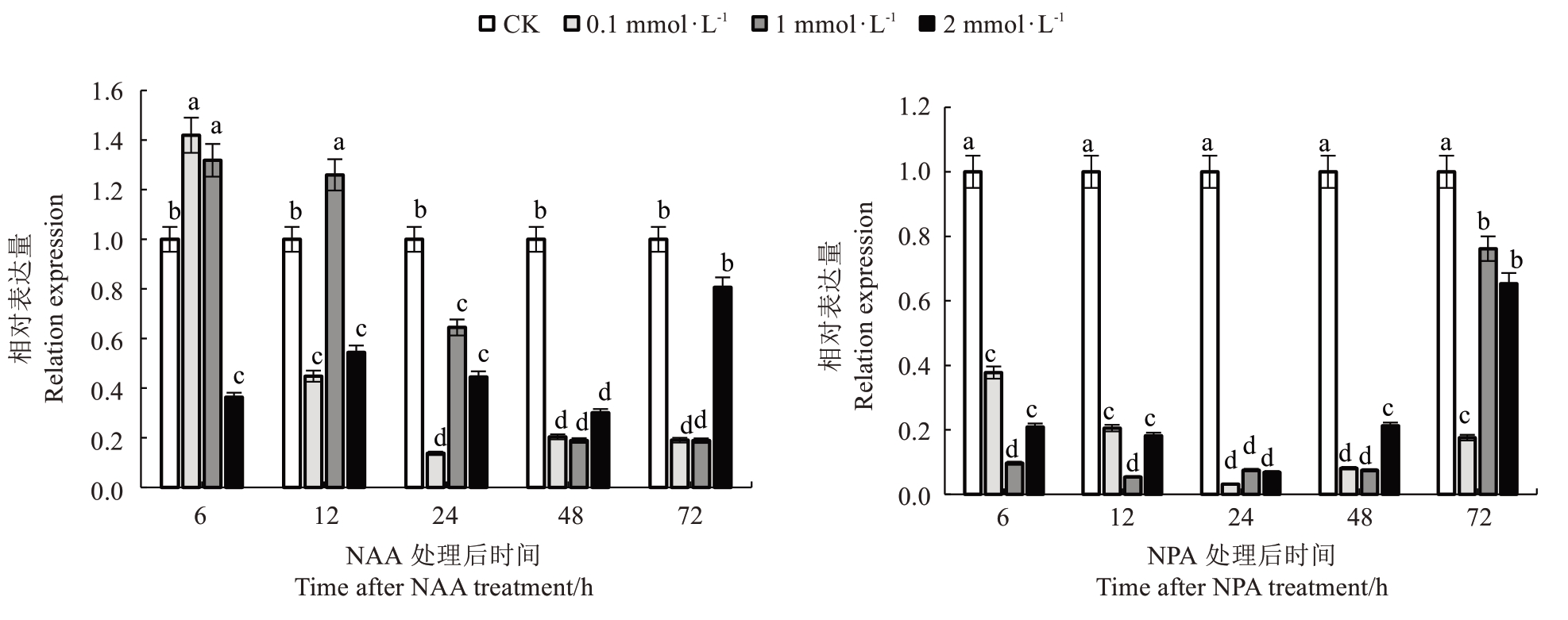
图3 不同浓度NAA 和NPA 处理后不同时间PpIAA11 的表达分析
Fig. 3 Analysis of PpIAA11 expression at different time after treatment with different concentrations of NAA and NPA
2.5 转基因番茄植株的鉴定及过表达PpIAA11 对果实生长发育的影响
为探究PpIAA11 在桃果实发育中的功能,对T1代转基因番茄植株进行分子鉴定。提取野生型与转基因株系叶片的总DNA 进行PCR 检测,结果显示10 个转基因株系能扩增出PpIAA11 目的条带,而野生型中则无相应扩增产物。qRT-PCR 分析进一步表明,10 个转基因性株系中的PpIAA11 相对表达量均显著高于野生型,其中T1_2 和T1_3 株系的表达水平最高(图4)。因此,后续试验选择T1_2 和T1_3 作为代表性高表达株系,用于进一步的功能分析。
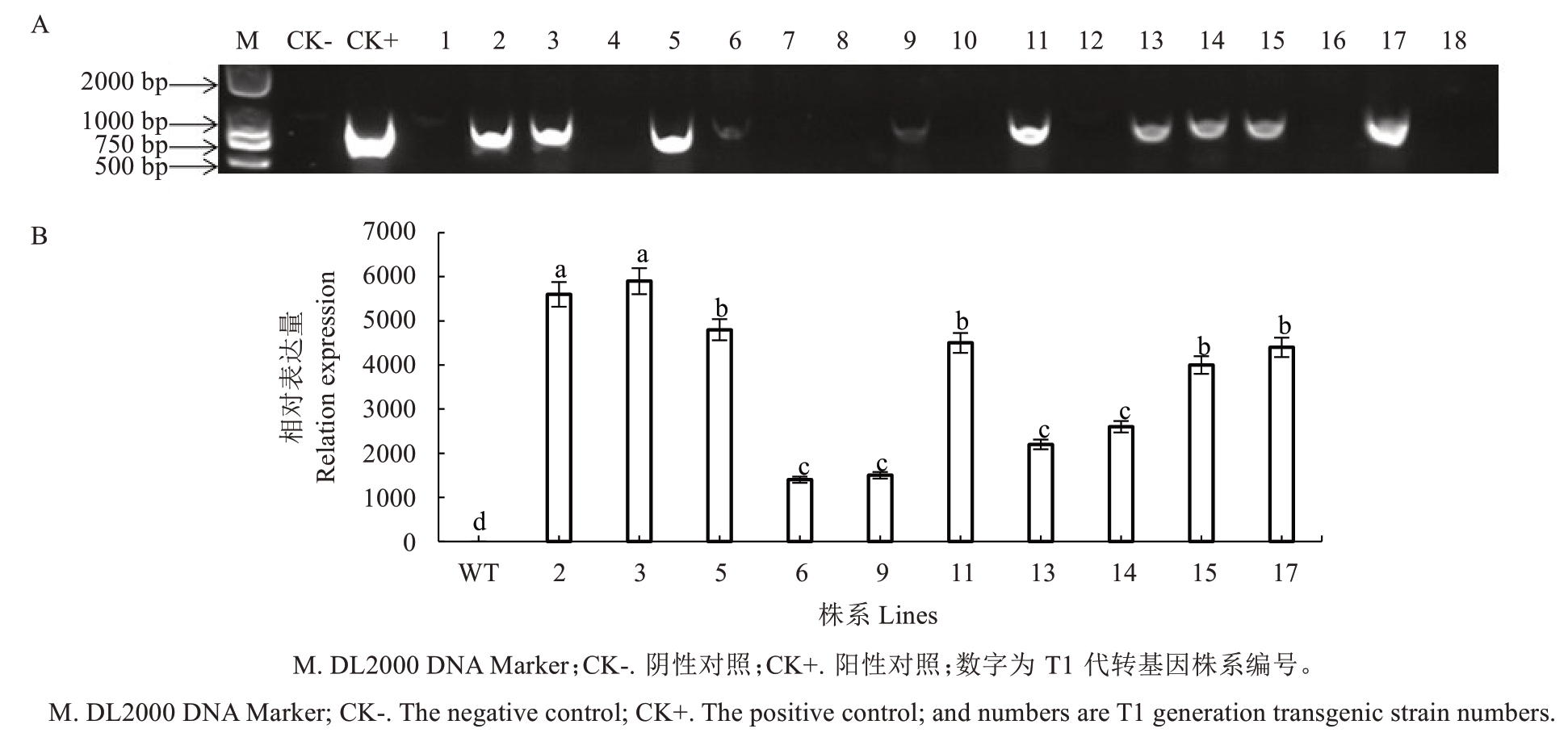
图4 转基因番茄阳性植株鉴定(A)及表达分析(B)
Fig. 4 Identification (A) and expression analysis (B) of transgenic tomato positive plants
经观察发现,与野生型番茄相比,PpIAA11 过表达株系(T1_2和T1_3)的果实形态发生显著变化,即果实具有明显的果尖(图5-A)。经测定,转基因株系果实的果尖长度2~3 mm,而野生型果实则未观察到该表型(图5-B)。此外,从开花至果实成熟,T1_2和T1_3 株系的果实成熟所需时间平均比野生型缩短约5 d(图5-C)。以上结果表明,PpIAA11 可能通过介导生长素信号通路,在调控果实形态建成和成熟时序中发挥重要作用。
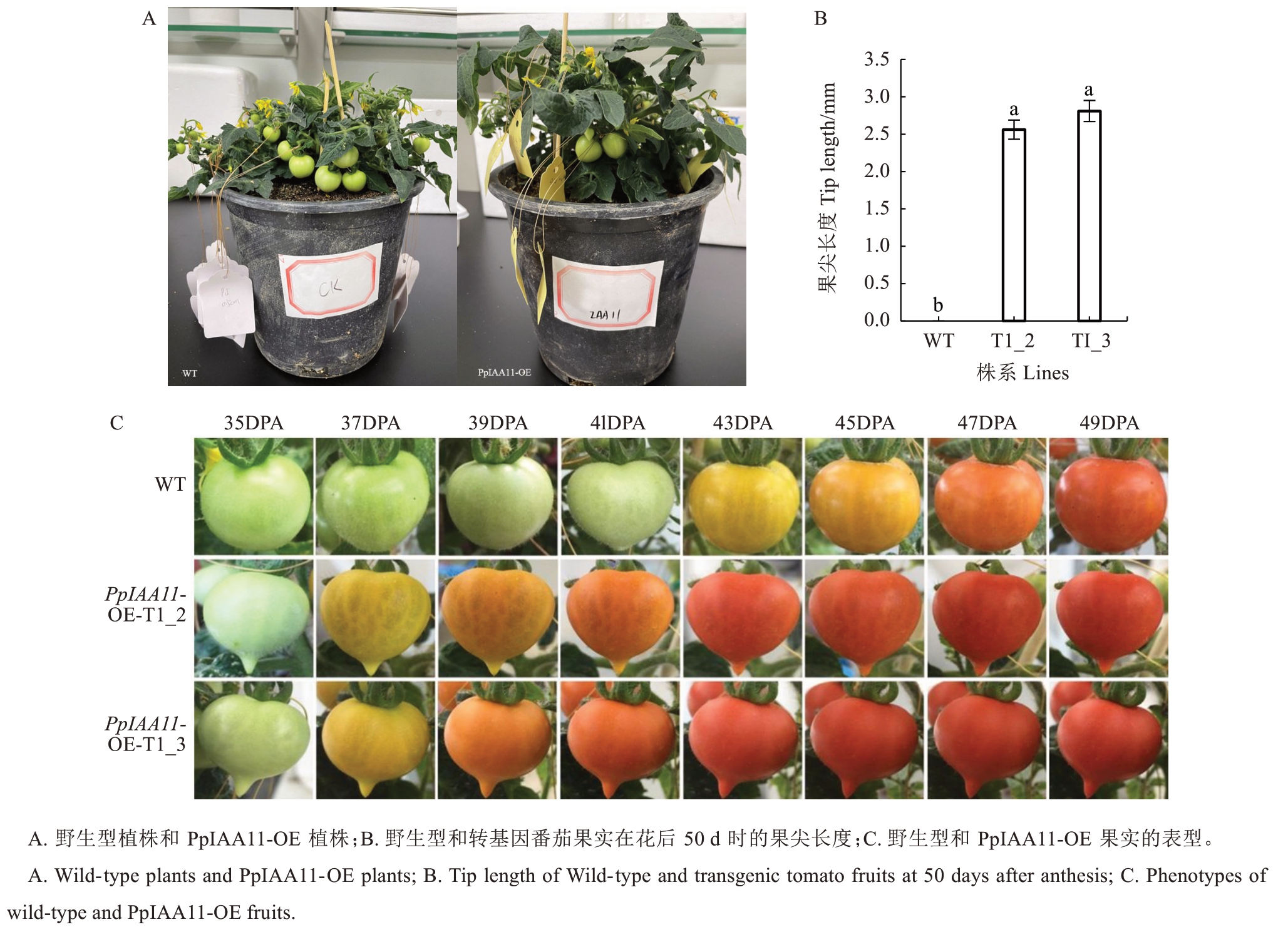
图5 转基因番茄植株形态、果尖长度及果实表型
Fig. 5 Transgenic tomato plant morphology, fruit tip length and fruit phenotype
3 讨 论
生长素作为关键植物激素,广泛参与胚胎发育、花器官形成、果实发育与成熟等多个生物学过程,并通过复杂的信号转导网络调控植物对环境的适应性反应[29]。在生长素信号通路中,AUX/IAA 家族蛋白作为一类早期响应因子和抑制因子,通过蛋白降解介导的生长素信号传递,在维持植物体内激素动态平衡中发挥核心作用[30]。桃基因组中已鉴定出多个AUX/IAA 家族成员,其中部分基因已被证实参与调控果实发育与成熟进程[31-33]。笔者聚焦于PpIAA11,通过多维度试验解析了其在桃果实发育过程中的功能及调控机制。本团队对前期转录组数据分析发现,PpIAA11在日川白凤桃果实的第2次快速膨大期表达量显著上调[25]。笔者进一步通过分子克隆获得CDS 序列,全长939 bp,编码312 个氨基酸。进化分析表明PpIAA11 与蔷薇科近缘种(如巴旦木PdIAA11 和欧洲甜樱桃PaIAA11)具有高度序列相似性和保守的结构域特征,表明其在蔷薇科果树中具有一定功能保守性[33]。然而,不同桃品种中PpIAA11的表达模式存在明显差异。在日川白凤中,其表达量在第2次快速膨大期达到峰值,而在晚24号中,则在成熟后期表达最高[15]。这一现象表明,PpIAA11可能在不同遗传背景下受到差异调控,其功能或许具有品种特异性,这为今后开展桃品种改良的分子育种提供了重要线索。
启动子顺式元件分析发现,PpIAA11 启动子区富含激素响应元件,尤其是ABA(9 个)和MeJA(7个)响应元件,远多于生长素响应元件(2 个)。这一结果表明,PpIAA11 的表达可能更易受到ABA 和茉莉酸途径的调控,而非直接受生长素诱导。这为解析AUX/IAA 基因家族在不同激素交叉调控中的复杂调控模式提供了新视角。在PpIAA11 对外源生长素及其抑制剂的响应方面,观察到PpIAA11 对NAA和NPA 处理的浓度和时间具有双重依赖性。低浓度(0.1 mmol·L-1)NAA 和NPA 处理能引起PpIAA11表达的显著变化,并在24 h趋于稳定,这与草莓FvIAA17的响应模式相似[34];而高浓度(1.0、2.0 mmol·L-1)NPA 处理下,PpIAA11 表达变化缺乏规律性,反映出生长素信号响应中存在复杂的浓度阈值效应和反馈调节机制。这一结果与植物激素在果实品质形成中具有浓度效应差异性的结论相吻合[23],也提示笔者在今后研究中,需更加注重激素处理浓度与时效之间的关系。
为深入解析PpIAA11 的生物学功能,通过在番茄中异源过表达该基因,发现其不仅使果实提前约5 d 成熟,还可以诱导形成明显的果尖,这一表型与前期在桃中PpIAA19 的研究结果类似[12]。这表明PpIAA11 可能通过影响生长素信号转导过程,改变果形发育进程,并促进果实乙烯的释放,从而加速果实成熟。然而,与本研究中PpIAA11 促进成熟的功能相反,同一家族中PpIAA5被报道可延缓桃果实成熟[33]。以上结果表明,桃AUX/IAA 基因家族内部可能存在功能的重叠和分化,不同成员可能通过调控生长素信号通路中相同或不同下游靶基因,也可能与相同或不同的ARF 转录因子互作,从而在果实成熟过程中扮演相同、不同甚至相反的调控角色。笔者初步揭示了PpIAA11在果实成熟过程中的潜在重要作用。然而,由于番茄与桃的果实类型和调控机制存在差异,PpIAA11 基因在桃果实成熟的信号通路中具体与哪些转录因子互作,以及如何调控下游基因的表达,仍需通过进一步的试验加以验证。
4 结 论
笔者成功克隆了桃生长素早期响应基因PpIAA11,并在番茄中进行了异源功能验证。结果表明,PpIAA11 的表达受外源生长素处理浓度和时间的动态调控,在番茄中过表达PpIAA11 可促进果实成熟并诱导果尖形成,这为解析桃果实发育过程中激素介导的形态建成和成熟调控机制提供了理论依据。
[1] 张雪,刘志民,陈华君,马焕普. 桃果实缝合线软化过程中内源激素的变化[J]. 果树学报,2008,25(2):172-177.ZHANG Xue,LIU Zhimin,CHEN Huajun,MA Huanpu. Changes of endogenous hormones during suture softening of peach fruit[J]. Journal of Fruit Science,2008,25(2):172-177.
[2] 曾文芳,王志强,潘磊,刘慧,牛良,鲁振华,崔国朝. 生长素对油桃‘24-30’果实软化和乙烯生物合成的影响[J]. 果树学报,2015,32(2):200-205.ZENG Wenfang,WANG Zhiqiang,PAN Lei,LIU Hui,NIU Liang,LU Zhenhua,CUI Guochao. Effect of NAA on fruit softening and ethylene biosynthesis of ‘24-30’ nectarine[J]. Journal of Fruit Science,2015,32(2):200-205.
[3] 李阳,张巍,杨爱珍,刘悦萍. 桃果实发育期间内源IAA 含量变化[J]. 北京农学院学报,2011,26(4):4-6.LI Yang,ZHANG Wei,YANG Aizhen,LIU Yueping. The content change of IAA during peach fruits development[J]. Journal of Beijing University of Agriculture,2011,26(4):4-6.
[4] MOCKAITIS K,ESTELLE M. Auxin receptors and plant development:A new signaling paradigm[J]. Annual Review of Cell and Developmental Biology,2008,24:55-80.
[5] DHARMASIRI N,DHARMASIRI S,ESTELLE M. The F-box protein TIR1 is an auxin receptor[J]. Nature,2005,435(7041):441-445.
[6] CHAPMAN E J,ESTELLE M. Mechanism of auxin-regulated gene expression in plants[J]. Annual Review of Genetics,2009,43:265-285.
[7] GUILFOYLE T J. The PB1 domain in auxin response factor and Aux/IAA proteins:A versatile protein interaction module in the auxin response[J]. The Plant Cell,2015,27(1):33-43.
[8] LV B S,YU Q Q,LIU J J,WEN X J,YAN Z W,HU K Q,LI H B,KONG X P,LI C L,TIAN H Y,DE SMET I,ZHANG X S,DING Z J. Non-canonical AUX/IAA protein IAA33 competes with canonical AUX/IAA repressor IAA5 to negatively regulate auxin signaling[J]. The EMBO Journal,2019,39(1):e101515.
[9] OVERVOORDE P J,OKUSHIMA Y,ALONSO J M,CHAN A,CHANG C,ECKER J R,HUGHES B,LIU A,ONODERA C,QUACH H,SMITH A,YU G X,THEOLOGIS A. Functional genomic analysis of the AUXIN/INDOLE-3-ACETIC ACID gene family members in Arabidopsis thaliana[J]. The Plant Cell,2005,17(12):3282-3300.
[10] SU Y L,HE H H,WANG P,MA Z H,MAO J,CHEN B H. Genome-wide characterization and expression analyses of the auxin/indole-3-acetic acid (Aux/IAA) gene family in apple (Malus domestica) [J]. Gene,2021,768:145302.
[11] JING X T,ZOU Q,YANG H. Genome-wide identification and characterization of the Aux/IAA gene family in strawberry species[J]. Plants,2024,13(20):2940.
[12] DING Y F,ZENG W F,WANG X B,WANG Y,NIU L,PAN L,LU Z H,CUI G C,LI G H,WANG Z Q. Over-expression of peach PpIAA19 in tomato alters plant growth,parthenocarpy,and fruit shape[J]. Journal of Plant Growth Regulation,2019,38(1):103-112.
[13] HOU Y M,LI H X,ZHAI L L,XIE X,LI X Y,BIAN S M. Identification and functional characterization of the Aux/IAA gene VcIAA27 in blueberry[J]. Plant Signaling & Behavior,2020,15(1):e1700327.
[14] GUAN D,HU X,DIAO D H,WANG F,LIU Y P. Genomewide analysis and identification of the Aux/IAA gene family in peach[J]. International Journal of Molecular Sciences,2019,20(19):4703.
[15] 王巍. 生长素应答基因PpIAA5-ARF8 对桃果实成熟软化的调控作用[D]. 北京:北京农学院,2021.WANG Wei. Auxin-responsive gene PpIAA5-ARF8 on peach fruit ripening and softening[D]. Beijing:Beijing University of Agriculture,2021.
[16] LI J Y,TAO X Y,BU J W,YING T J,MAO L C,LUO Z S.Global transcriptome profiling analysis of ethylene-auxin interaction during tomato fruit ripening[J]. Postharvest Biology and Technology,2017,130:28-38.
[17] KUMAR R,AGARWAL P,PAREEK A,TYAGI A K,SHARMA A K. Genomic survey,gene expression,and interaction analysis suggest diverse roles of ARF and Aux/IAA proteins in Solanaceae[J]. Plant Molecular Biology Reporter,2015,33(5):1552-1572.
[18] HAMANN T,BENKOVA E,BÄURLE I,KIENTZ M,JÜRGENS G. The Arabidopsis BODENLOS gene encodes an auxin response protein inhibiting MONOPTEROS-mediated embryo patterning[J]. Genes & Development,2002,16(13):1610-1615.
[19] PURGATTO E,DO NASCIMENTO J R O,LAJOLO F M,CORDENUNSI B R. The onset of starch degradation during banana ripening is concomitant to changes in the content of free and conjugated forms of indole-3-acetic acid[J]. Journal of Plant Physiology,2002,159(10):1105-1111.
[20] LI X W,LIU P,ZHOU J Y,SU M S,MA Y P,JIA H J,DU J H,GAO Z S,YE Z W. Effects of exogenous application of GA4+7 and NAA on sugar accumulation and related gene expression in peach fruits during developing and ripening stages[J]. Journal of Plant Growth Regulation,2021,40(3):962-973.
[21] TADIELLO A,ZIOSI V,NEGRI A S,NOFERINI M,FIORI G,BUSATTO N,ESPEN L,COSTA G,TRAINOTTI L. On the role of ethylene,auxin and a GOLVEN-like peptide hormone in the regulation of peach ripening[J]. BMC Plant Biology,2016,16(1):44.
[22] LOHANI S,TRIVEDI P K,NATH P. Changes in activities of cell wall hydrolases during ethylene-induced ripening in banana:Effect of 1-MCP,ABA and IAA[J]. Postharvest Biology and Technology,2004,31(2):119-126.
[23] YUE P T,LU Q,LIU Z,LV T X,LI X Y,BU H D,LIU W T,XU Y X,YUAN H,WANG A D. Auxin-activated MdARF5 induces the expression of ethylene biosynthetic genes to initiate apple fruit ripening[J]. New Phytologist,2020,226(6):1781-1795.
[24] TATSUKI M,NAKAJIMA N,FUJII H,SHIMADA T,NAKANO M,HAYASHI K I,HAYAMA H,YOSHIOKA H,NAKAMURA Y. Increased levels of IAA are required for system 2 ethylene synthesis causing fruit softening in peach (Prunus persica L. Batsch)[J]. Journal of Experimental Botany,2013,64(4):1049-1059.
[25] ZHANG Y P,SU Z W,LUO L J,WANG P K,ZHU X D,LIU J C,WANG C. Exogenous auxin regulates the growth and development of peach fruit at the expansion stage by mediating multiple-hormone signaling[J]. BMC Plant Biology,2023,23(1):499.
[26] HU W,FAGUNDEZ S,KATIN-GRAZZINI L,LI Y J,LI W,CHEN Y N,WANG X M,DENG Z N,XIE S X,MCAVOY R J,LI Y. Endogenous auxin and its manipulation influence in vitro shoot organogenesis of citrus epicotyl explants[J]. Horticulture Research,2017,4:17071.
[27] CAO M,CHEN R,LI P,YU Y Q,ZHENG R,GE D F,ZHENG W,WANG X H,GU Y T,GELOVÁ Z,FRIML J,ZHANG H,LIU R Y,HE J,XU T D. TMK1-mediated auxin signalling regulates differential growth of the apical hook[J]. Nature,2019,568(7751):240-243.
[28] 程杰,张新圣,李安琪,姜晶. 番茄果实成熟过程中SlSWEET7a 的功能分析[J]. 中国农业科学,2018,51(15):2958-2968.CHENG Jie,ZHANG Xinsheng,LI Anqi,JIANG Jing. Functional analysis of SlSWEET7a gene during maturation of tomato fruits[J]. Scientia Agricultura Sinica,2018,51(15):2958-2968.
[29] BENJAMINS R,SCHERES B. Auxin:The looping star in plant development[J]. Annual Review of Plant Biology,2008,59:443-465.
[30] HOU Q D,QIAO G,SHANG C Q,SHEN L N,ZHOU K,MIN Y,WEN X P. Expression analyses of sweet cherry AUX/IAAs and involvements of PavAUX/IAA9/11 in regulating fruitlet abscission[J]. Plant Cell Reports,2025,44(7):143.
[31] SU L Y,BASSA C,AUDRAN C,MILA I,CHENICLET C,CHEVALIER C,BOUZAYEN M,ROUSTAN J P,CHERVIN C. The auxin Sl-IAA17 transcriptional repressor controls fruit size via the regulation of endoreduplication-related cell expansion[J]. Plant & Cell Physiology,2014,55(11):1969-1976.
[32] WANG X B,PAN L,WANG Y,MENG J R,DENG L,NIU L,LIU H,DING Y F,YAO J L,NIEUWENHUIZEN N J,AMPOMAH-DWAMENA C,LU Z H,CUI G C,WANG Z Q,ZENG W F. PpIAA1 and PpERF4 form a positive feedback loop to regulate peach fruit ripening by integrating auxin and ethylene signals[J]. Plant Science,2021,313:111084.
[33] QIN Y F,WANG W,CHANG M M,YANG H Q,YIN F R,LIU Y P. The PpIAA5-ARF8 module regulates fruit ripening and softening in peach[J]. Horticulturae,2023,9(10):1149.
[34] TRAINOTTI L,TADIELLO A,CASADORO G. The involvement of auxin in the ripening of climacteric fruits comes of age:The hormone plays a role of its own and has an intense interplay with ethylene in ripening peaches[J]. Journal of Experimental Botany,2007,58(12):3299-3308.